Original data
In this example, we use the data presented in Johannesen et al. and available for download. It comprises the 4 compounds dofetilide, quinidine, ranolazine, and verapamil. The clinical 5-way crossover study administered single doses of 500 µg dofetilide, 400 mg quinidine sulfate, 1500 mg ranolazine, 120 mg verapamil hydrochloride, or placebo to 22 healthy subjects. PK and QT data were collected at 16 matching time points over 24 h after dose administration.
For this example we will use only the placebo and quinidine periods. The following R code was used to read the original excel dataset, filter it to keep only the placebo and quinidine periods and save it as csv.
data <- openxlsx::read.xlsx("SCR002_clinicalData4drugs_originalFile.xlsx")
Quinidine_data <- data %>% filter(EXTRT=="Placebo" | EXTRT=="Quinidine Sulph") %>% select(-c(ARMCD))
write.csv(Quinidine_data, file = "Quinidine_data.csv", quote=F, row.names = F, na=".")
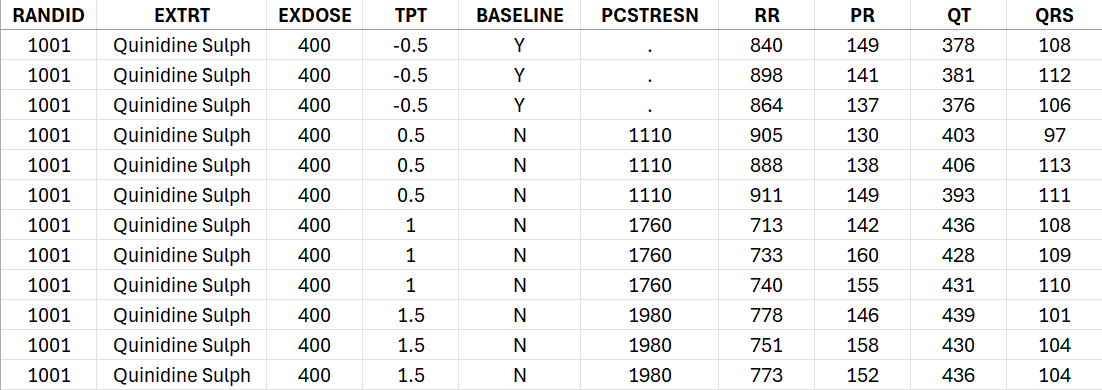
The csv file contains the following columns:
Data processing
In order to use the provided R functions, the R script mlxQTcTools.R must be sourced. In addition, a monolixSuite installation must be available and the lixoftConnectors R package installed. In order to indicate the location of the MonolixSuite installation, the functioninitQTc(path=...) is used.
source('../_mlxQTcTools/mlxQTcTools.R')
initQTc(path="C:/Program Files/Lixoft/MonolixSuite2024R1")
In order to perform a concentration-QTc analysis, it is necessary to apply the following processing:
-
calculate heart-rate correction for QT to generate QTc, for instance using Fridericia’s formula
-
average triplicate QTc for each time point
-
calculate the baseline-corrected QTc, i.e ΔQTc. In our case, the baseline is simply the pre-dose measument at time -0.5 hr. It is indicated in the dataset with BASELINE=Y
-
given that both placebo and quinidine data is available for each individual, it is also possible to calculate directly the baseline-corrected and placebo-corrected QTC, i.e ΔΔQTc
-
for exploratory data visualization purpose, the ΔHR and ΔΔHR can also be computed
Covariates and regressor variables which will appear in the conc-QTc model are also added as additional columns:
-
the centered QTc baseline, and placebo-adjusted centered baseline in case of ΔΔQTc
-
time as categorical factor
-
concentration and RR interval duration
The steps can be performed using the function process_QTcData():
process_QTcData(data="Quinidine_data.csv",
QTname = "QT", RRname = "RR", CONCname = "PCSTRESN",
TRTname = "EXTRT", IDname = "RANDID", TIMEname = "TPT",
bComputeQTc = TRUE, correctionMethod = "Fridericia",
bComputeBaseline = TRUE, BLFLAGname = "BASELINE", baselineType = "predose",
stepOrder = "FirstCorrectThenAverage",
outName = "Quinidine_formatted.csv", silent=F)
The column containing the QT, RR, drug concentration, treatment (placebo or drug), subject id and time are indicated using the arguments QTname, RRname, CONCname, TRTname, IDname, TIMEname. Note that the values present in the TRTname column will be used as plot labels for the report. In this example, the column contains “Placebo” and “Quinidine Sulph”. If several dose levels are used, the TRTname column should contain different values for the different dose levels, for instance “Placebo”, “Quinidine Sulph 250 ug” and “Quinidine Sulph 500 ug”.
In order to use only a subset of the categories available in the TRTname column, it possible to list the categories to include in the optional TRT argument. If the category corresponding to placebo is different from “placebo”, the name of the placebo category should be specified in the placeboNameargument.
If the QTc column (heart rate-corrected QT) is not present in the dataset, it can be computed using bComputeQTc = TRUE. Several correction methods are available and can be chosen using correctionMethod = "Fridericia", "Bazett", "Framingham", "Hodges" or “Population”. If the QTc column is already present, it should be indicated using the QTCname argument.
When several replicates per time points are present, they will be averaged. It is possible to first average triplicate and then apply the heart rate correction or do the opposite using stepOrder = "FirstCorrectThenAverage" or "FirstAverageThenCorrect". Records are grouped by TIME and TRT value. If another column should be used for grouping (e.g DAY), use the addVariables argument.
If the QTc baseline is not present yet in the dataset, it can be computed by giving bComputeBaseline = TRUE and indicating the baseline flag column in BLFLAGname. The baseline flag column should contain “0”/”1” values or “N”/”Y” values with “Y” and “1” indicating that baseline records. If several baseline records are available for each period (i.e treatment) of each individual, the baseline values will be averaged. If the baseline is already computed, the arguments BLQTCname (QTc baseline) and BLHRname (HR baseline) must be provided. The baseline type (“predose” or “time-matched”) is given with the baselineType argument. If the baseline should be computed using the addVariables for grouping, use bComputeBLperAddVar=T.
The name of the output file is given via the outName argument.
Running the function on the Dofetilide example generates the following console output:
Warning: The column 'QT' contains non-numerical values. They will be replaced by NA.
Warning: The column 'PCSTRESN' contains non-numerical values. They will be replaced by NA.
Info: The following treatment groups have been found: Placebo, Quinidine Sulph
Info: The HR column has not been provided and has thus been calculated based on the 'RR' column.
Info: The missing concentrations in the placebo treatment have been replaced by zeros.
Info: The missing concentrations on rows with BASELINE=Y or 1 have been replaced by zeros.
Info: The QTc column has been calculated based on the 'QT' and 'RR' columns, using Fridericia's formula: QTc=QT/(RR/1000)^(1/3)
Info: The lines corresponding to the same id, same time and same treatment (e.g triplicates) have been averaged.
Info: The BLQTc and BLHR columns have been calculated for each combination of RANDID, EXTRT, using the lines flagged as BASELINE=Y or 1.
Info: The dQTc and dHR columns have been calculated as dQTc=QTc-BLQTc and dHR=HR-BLHR.
Info: The BLQTc_cent column has been calculated as BLQTc_cent=BLQTc-mean(BLQTc).
Info: The ddQTc, ddHR and BLQTc_centAdjPl columns have be computed by subtracting the placebo data.
Info: The Cc_reg (concentration as regressor), RR_reg (RR as regressor) and TIME_cat (TIME as categorical covariate) columns have been added.
==> Created new dataset Quinidine_formatted.csv.
The Info lines give details about the data processing steps. The Warning lines draw attention on unusual situations such as missing concentration on lines not belonging to the placebo group. The console output can be avoided using silent=T.
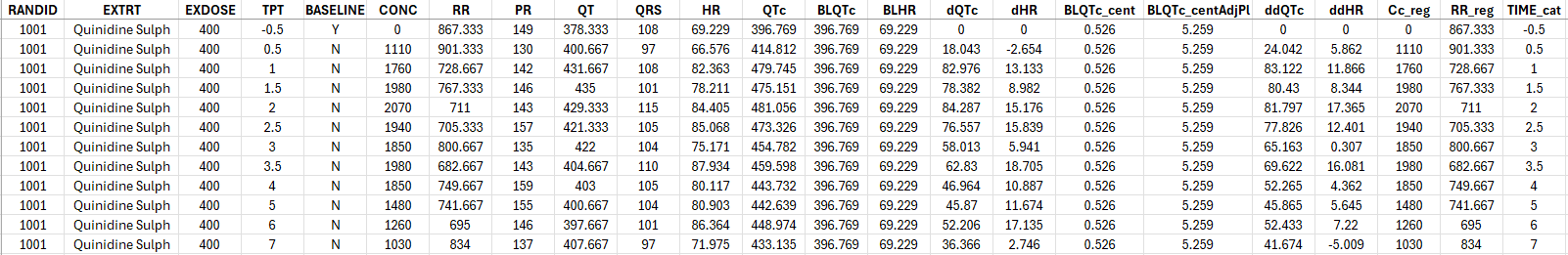
The generated formatted dataset is the following. It contain all columns needed to create a Monolix project for conc-QTc analysis as well as creating the usual exploratory data analysis plots.
Model fitting
The next step is to generate and run Monolix projects to estimate the parameters of the concentration-QTc models. Both linear and non-linear conc-QTc relationships are considered.
To generate and run the Monolix projects to run the con-QTc analysis, the function generateAllQTcProjects() is used. Given that each individual had both placebo and active drug data and ΔΔQTc could be calculated from the data, we will model ΔΔQTc directly.
generateAllQTcProjects(dataSet="Dofetilide_formatted.csv",
IDname="RANDID", TIMEname="TPT", TRTname="EXTRT", CONCname="CONC", placeboName = "Placebo",
modelFolder="../_mlxQTcTools/models/ddQTcModels",
projectFolder="./mlx_runs",
isddQTC = T,
bRun = T)
The user should provide the formatted dataset in the argument dataSet, as well as the name of the columns containing the subject id in IDname, time in TIMEname, treatment groups in TRTname, concentration in CONCname. The other columns (e.g QT, QTc, HR, etc) must have predefined headers and do not need to be specified.
The category of the treatment group column representing placebo can be indicated using placeboName = "Placebo". If this argument is omitted, the function will search for the string “placebo” among the categories present in the TRT column. If the dataset contains no placebo data, use placeboName = "no placebo".
The folder containing the structural models to be used to generate the monolix runs are given in the argument modelFolder. The R package contains a pre-written library of models with linear, Emax, Emax with sigmoidicity and loglinear conc-QTc relationship. User-defined models can be added to this folder is needed.
In folder in which the runs are generated is the projectFolder argument.
Whether conc-ΔΔQTc or conc-ΔQTc should be run is indicated using isddQTC = T or F. To run the projects after having created them, use bRun = T.
When running the function, the console output indicates the generated runs, the implemented models and the progress of the parameter estimation:
Created project ./mlx_runs/ddQTc_Emax.mlxtran with model:
ddQTc = theta1 + theta4 * BLQTc_centAdjPl + (theta2 + eta2) * Conc/(Conc + theta5)
Running project... Done.
Created project ./mlx_runs/ddQTc_EmaxSigma.mlxtran with model:
ddQTc = theta1 + theta4 * BLQTc_centAdjPl + (theta2 + eta2) * Conc^theta6/(Conc^theta6 + theta5^theta6)
Running project... Done.
Created project ./mlx_runs/ddQTc_Linear.mlxtran with model:
ddQTc = theta1 + theta4 * BLQTc_centAdjPl + (theta2 + eta2) * Conc
Running project... Done.
Created project ./mlx_runs/ddQTc_Linear_effectComp.mlxtran with model:
ddQTc = theta1 + theta4 * BLQTc_centAdjPl + (theta2 + eta2) * Ce with dCe/dt = (Conc - Ce)/theta5
Running project... Done.
Created project ./mlx_runs/ddQTc_LogLinear.mlxtran with model:
ddQTc = theta1 + theta4 * BLQTc_centAdjPl + (theta2 + eta2) * log(1 + Conc/theta5)
Running project... Done.
The generated Monolix runs can be opened with the Monolix GUI for inspection.
Report generation
A standard report can be generated using the function quarto_render() and a Quarto template. The quarto template will read the results from generated Monolix projects and generate a report including:
-
Exploratory data analysis (data summary, QT/QTc, heart rate correction, baseline QTc, ΔQTc, ΔΔQTc and concentration-time)
-
Model assumption assessments (heart-rate independence from drug concentration, QTc independence from heart rate, linearity of the concentration-QTc relationship, immediate effect of concentration change on ΔΔQTc change)
-
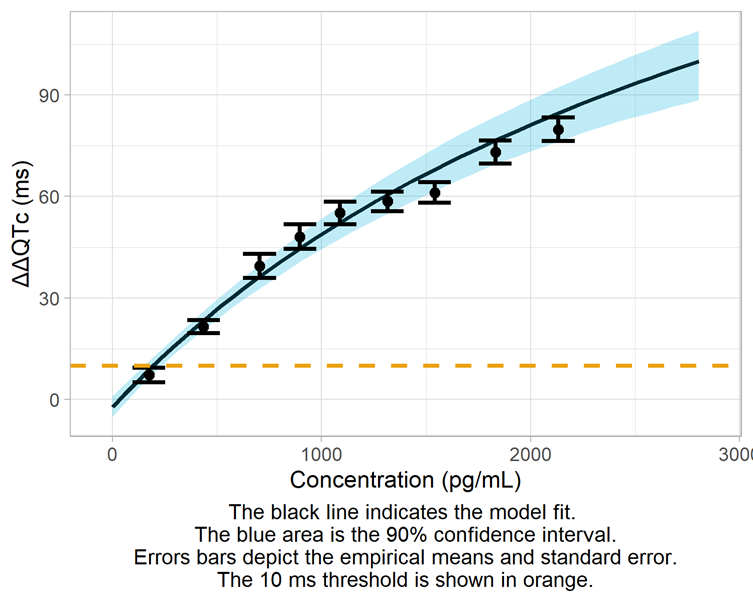
Modeling results (comparison of the different models, model fit, parameter estimates, goodness of fit)
-
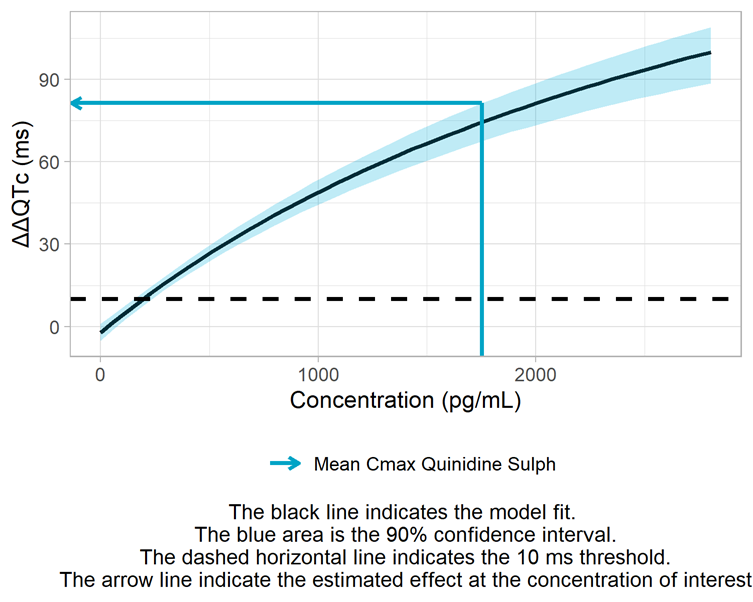
Derivation of the ΔΔQTc prediction intervals, including assessment of the 10-ms threshold
The quarto function is called in the following way:
quarto_render(input = "../_mlxQTcTools/QTc_quarto_allModels.qmd",
output_file = "Quinidine_ddQTc_report_new.docx",
execute_params = list(folderName = "../Example_Quinidine/mlx_runs", # give relative path from quarto template (.qmd) to folder containing runs
compoundName = "Quinidine",
TRTcolumn = "EXTRT",
STRATname = "Treatment",
bHasPlacebo = TRUE,
placeboName = "Placebo",
studyType = "ddQTc",
concentrationUnit = "pg/mL",
nBins = 10,
orderTRT = c("Placebo","Quinidine Sulph"),
thresBICc = 0))
The input is the quarto template. A standard quarto template is provided in the package but it can also be adjusted to the users need. The output_file indicates the name of the generated report as word document. The execute_params list the folder containing the Monolix runs in folderName, the string to be used as compound name in compoundName, the column of the formatted dataset containing the concentration in CONCcolumn, the column containing the treatment categories in TRTcolumn, the header/label to be used for the stratification column (here the TRTcolumn) in STRATname, a logical to indicate whether the dataset contains placebo data in bHasPlacebo, the placebo name in the treatment group column in placeboName, the type of study using studyType = "dQTc"or “ddQTc”, the concentration unit in concentrationUnit(for plot labels), the number of bins to average the concentration in nBins, the order of the categories present in the treatment group column to be used in plots and tables in orderTRT and the BICc threshold to consider that an alternative model is better than the linear model in thresBICc.
Note that the folderName must indicate the relative path from the .qmd template (not the R working directory) to the folder containing the runs.
The generated report can be visualized below:

Download the generated the ΔΔQTc quinidine report (word format).
Conclusion
The conc-ΔΔQTc data for Quinidine is best captured by an Emax model. The upper bound of the 90% confidence interval at the geometric mean maximum concentration is 81 ms, exceeding by far the 10 ms threshold.