[Monolix] Distribution of the individual parameters computed by Monolix
Plot the distribution of the individual parameters.
Usage
plotParametersDistribution(
parameters = NULL,
plot = "pdf",
settings = list(),
preferences = NULL,
stratify = list()
)
Arguments
parameters vector of parameters to display. (by default the first 4 computed parameters are displayed). plot (character) Type of plot: probability density distribution ("pdf"), cumulative density distribution ("cdf") (default "pdf) settings a list of optional plot settings:indivEstimate Calculation of individual estimates: conditional mean ("mean"), conditional mode with EBE's ("mode"), conditional distribution ("simulated") (default "simulated"). empirical (logical) If TRUE, plot empirical density distribution (default TRUE). theoretical (logical) If TRUE, plot theoretical density distribution (default TRUE). legend (logical) add (TRUE) / remove (FALSE) plot legend (default FALSE). grid (logical) add (TRUE) / remove (FALSE) plot grid (default TRUE). ncol (integer) number of columns when facet = TRUE (default 4). fontsize (integer) Plot text font size. preferences (optional) preferences for plot display, run getPlotPreferences("plotParametersDistribution") to check available displays. stratify List with the stratification arguments:groups - Definition of stratification groups. By default, stratification groups are already defined as one group for each category for categorical covariates, and two groups of equal number of individuals for continuous covariates. To redefine groups, for each covariate to redefine, specify a list with:namecharactercovariate name (e.g "AGE")definition(vector(continuous) || list>(categorical))For continuous covariates, vector of break values (e.g c(35, 65)). For categorical covariates, groups of categories as a list of vectors(e.g list(c("study101"), c("study201","study202"))) split (vector) - Vector of covariates used to split (i.e facet) the plot (by default no split is applied). For instance c("FORM","AGE"). filter (list< list> >) - List of pairs containing a covariate name and the vector of indexes or categories (for categorical covariates) of the groups to keep (by default no filtering is applied). For instance, list("AGE",c(1,3)) to keep the individuals belonging to the first and third age group, according to the definition in groups. For instance, list("FORM","ref") using the category name for categorical covariates.
Value
-
A ggplot object if one parameter,
-
A TableGrob object if multiple plots (output of grid.arrange)
See also
getChartsData getPlotPreferences
Examples
initializeLixoftConnectors(software="monolix")
project <- file.path(getDemoPath(), "1.creating_and_using_models",
"1.1.libraries_of_models", "theophylline_project.mlxtran")
loadProject(project)
runPopulationParameterEstimation()
runConditionalDistributionSampling()
plotParametersDistribution(parameters = "ka")
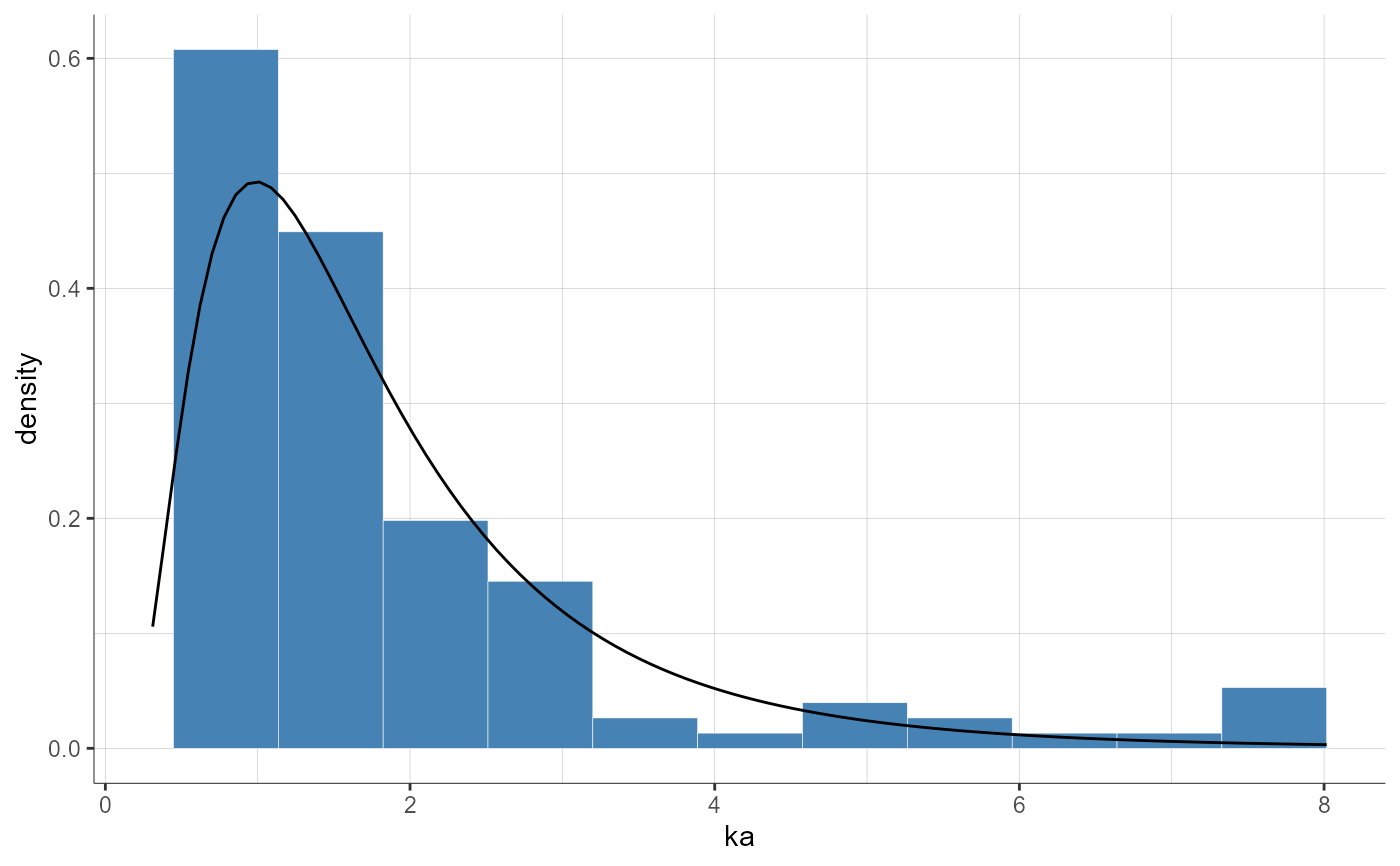
plotParametersDistribution(parameters = "Cl", plot = "pdf")
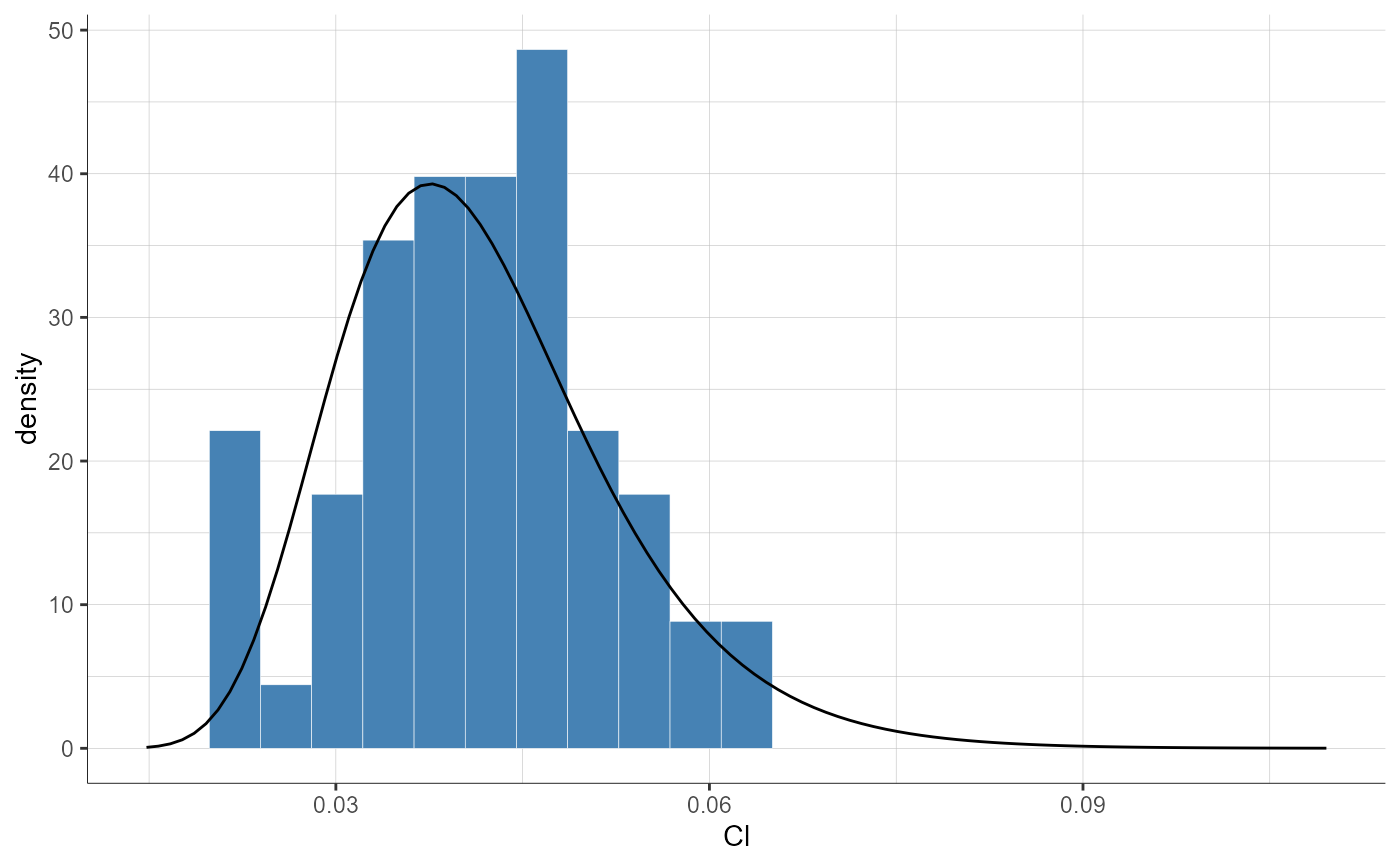
plotParametersDistribution(parameters = "ka", plot = "cdf")
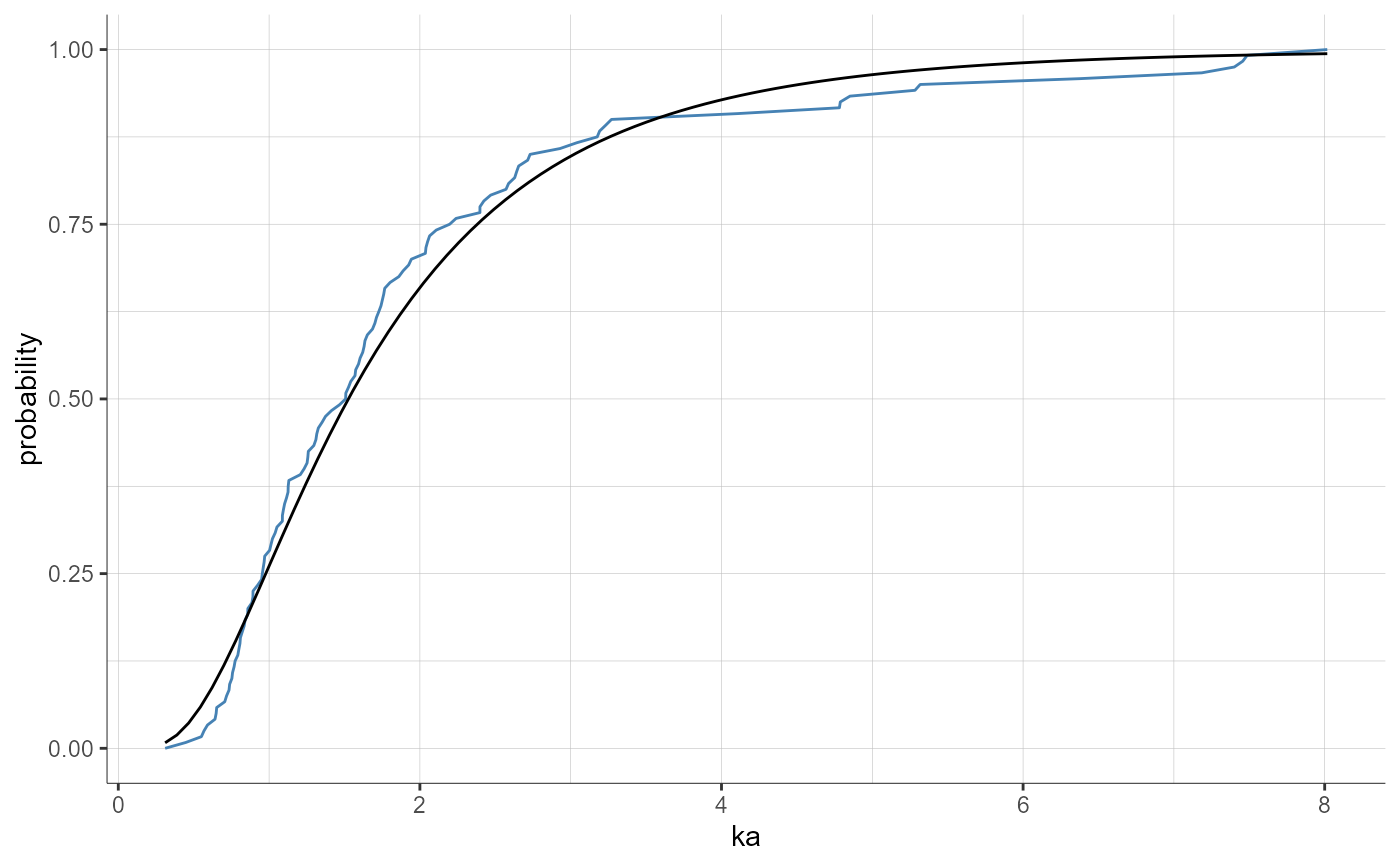
plotParametersDistribution(parameters = "ka", plot = "cdf",
settings = list(indivEstimate = "simulated"))
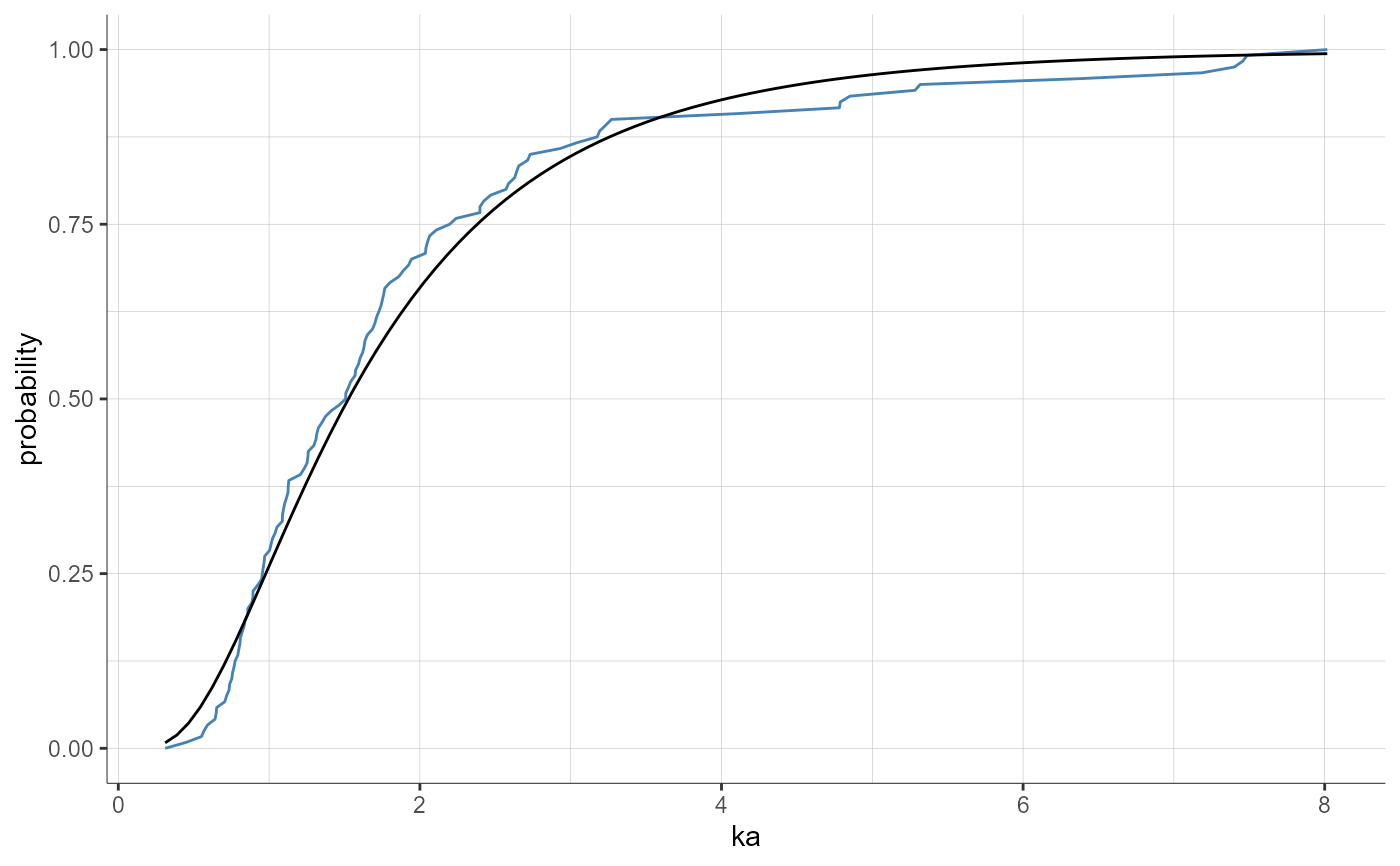
plotParametersDistribution(parameters = "Cl", plot = "pdf",
settings = list(theoretical = F))
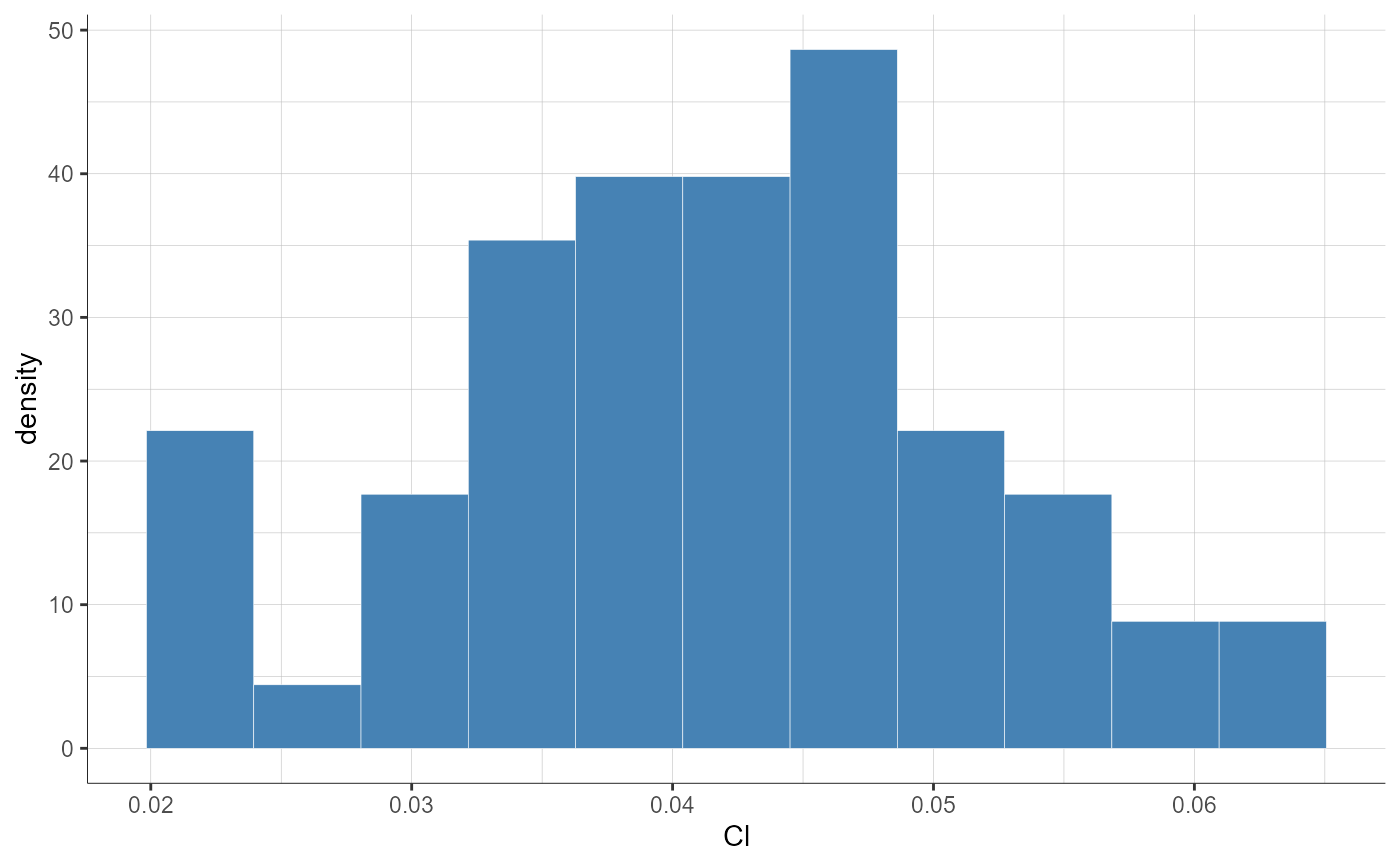
# stratification
plotParametersDistribution(stratify = list(filter = list("WEIGHT", 1),
groups = list(name = "WEIGHT", definition = 75)))
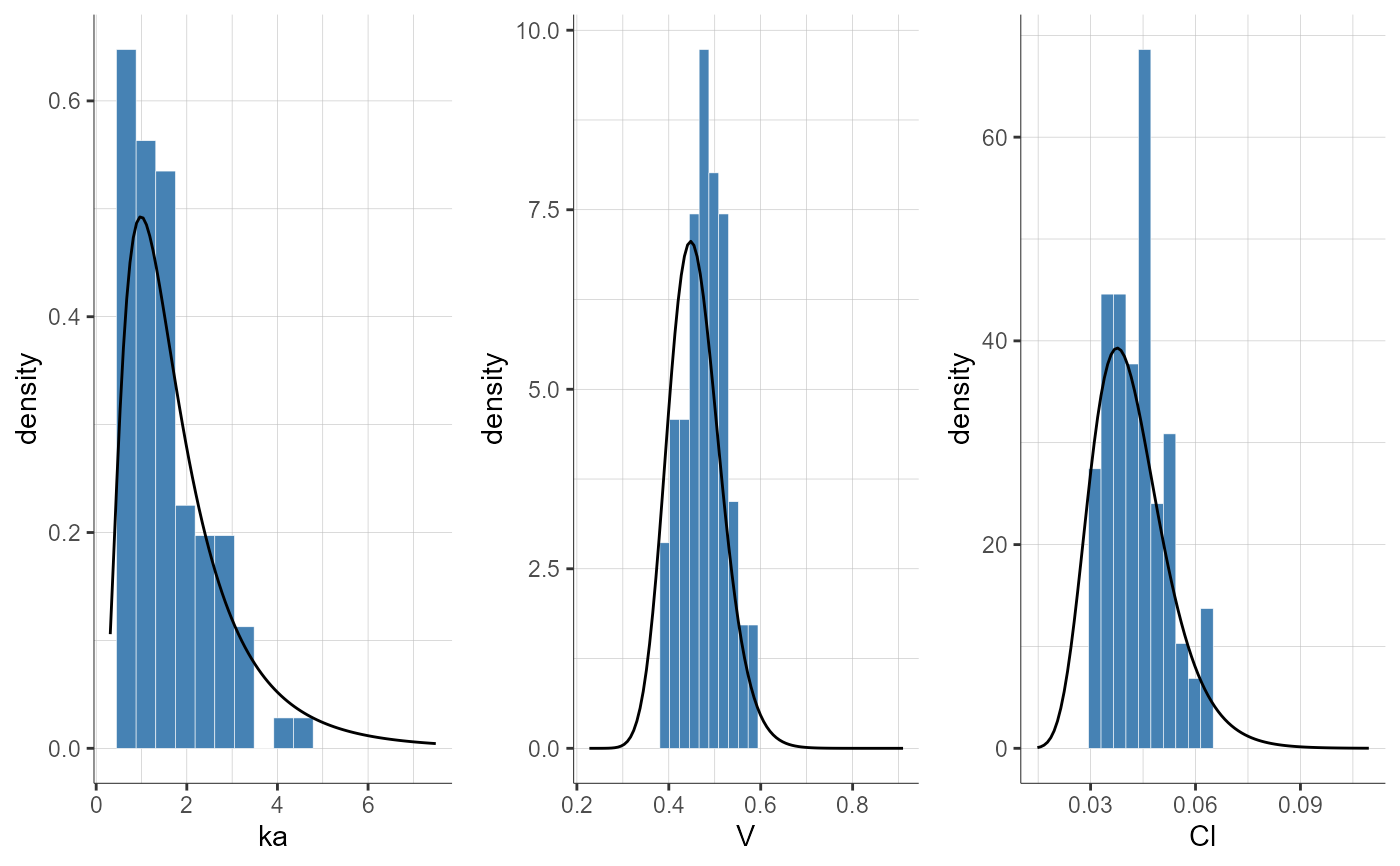
#> TableGrob (1 x 1) "arrange": 1 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[arrange]
plotParametersDistribution(parameters = "Cl", stratify = list(split = "SEX"))
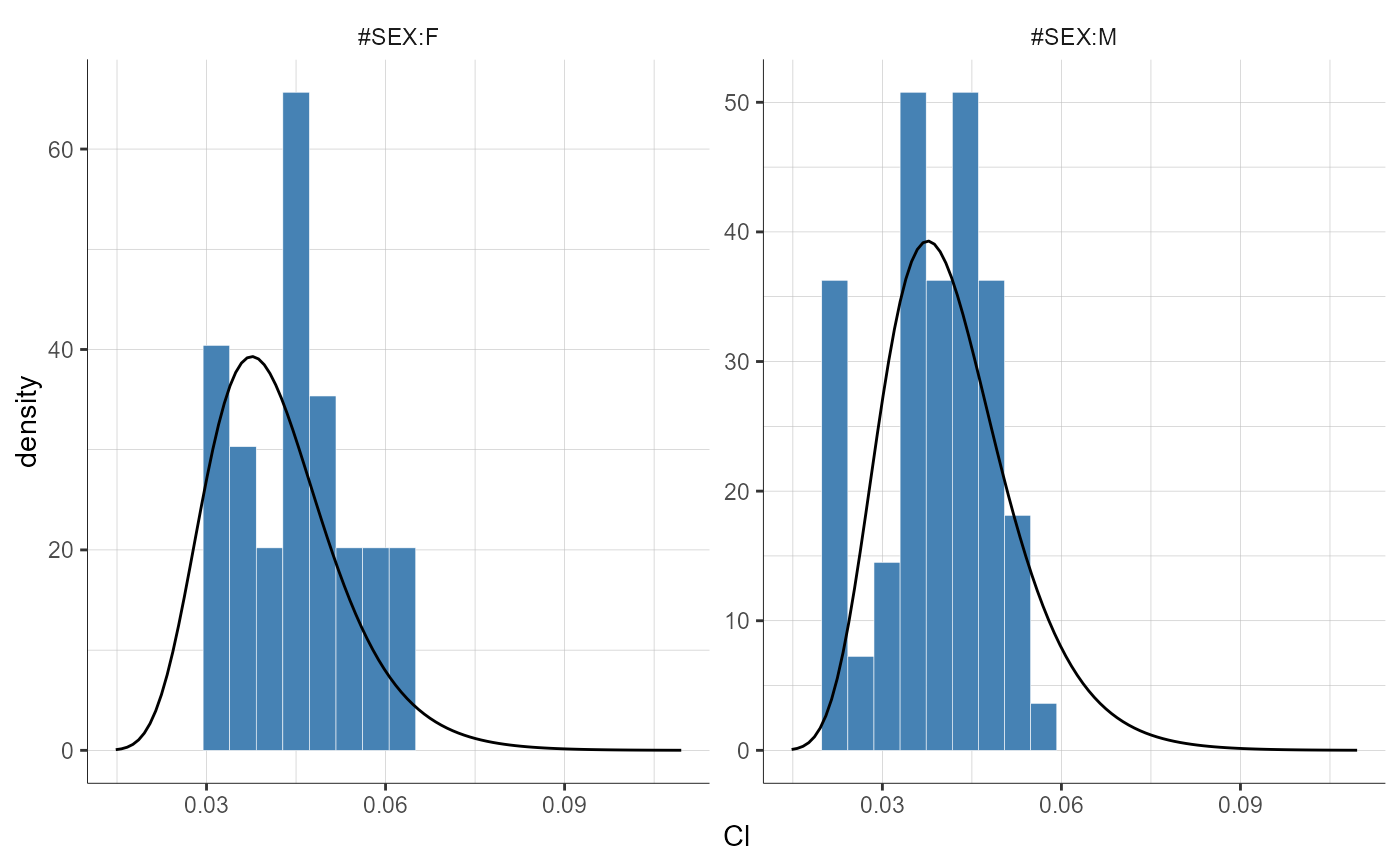
# update preferences
preferences = list(theoretical = list(color = "#B4468A", lineType = "solid", lineWidth = 0.8))
plotParametersDistribution(parameters = "ka", plot = "cdf", preferences = preferences)
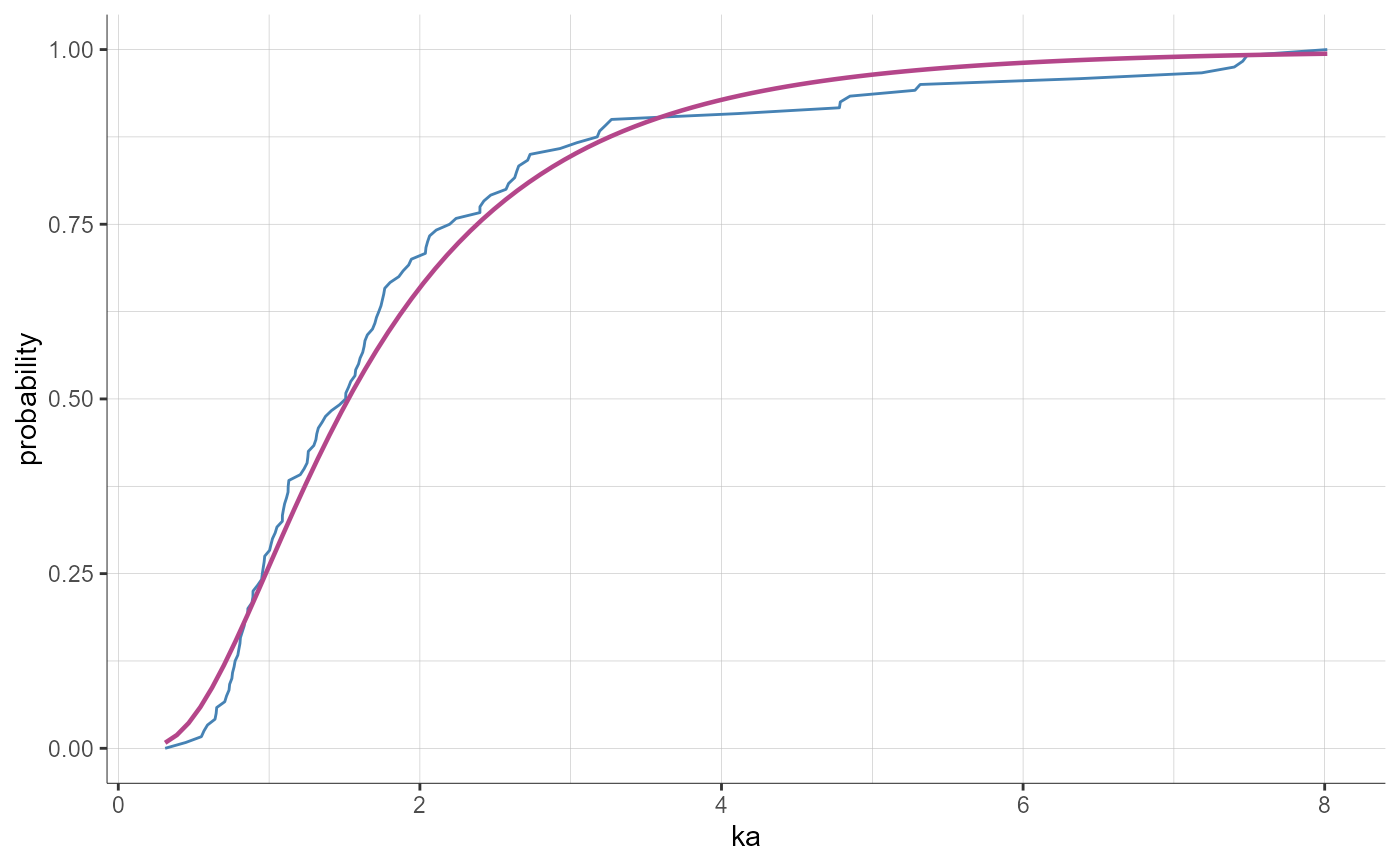
# multiple plots
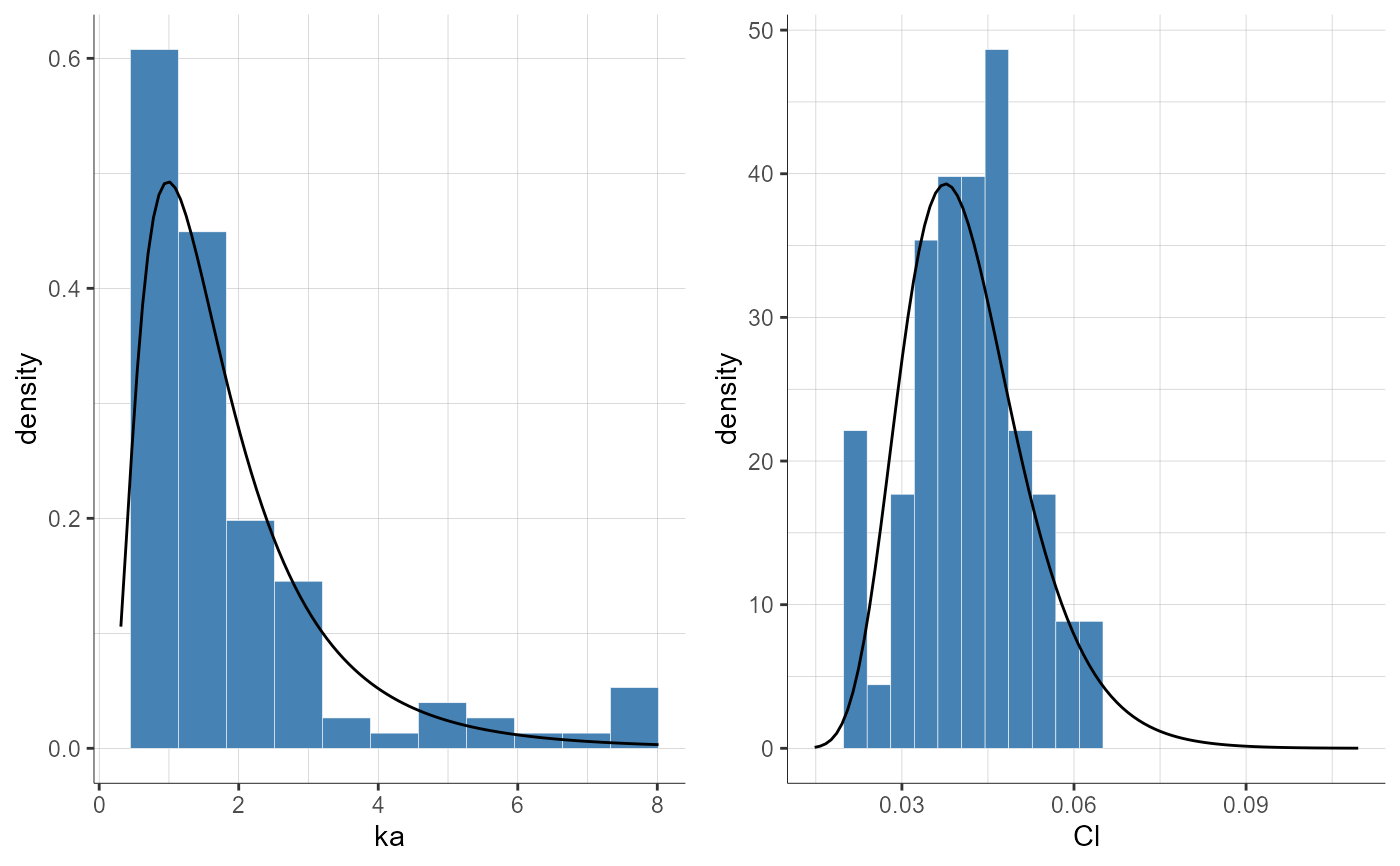
plotParametersDistribution(parameters = c("ka", "Cl"))
#> TableGrob (1 x 1) "arrange": 1 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[arrange]
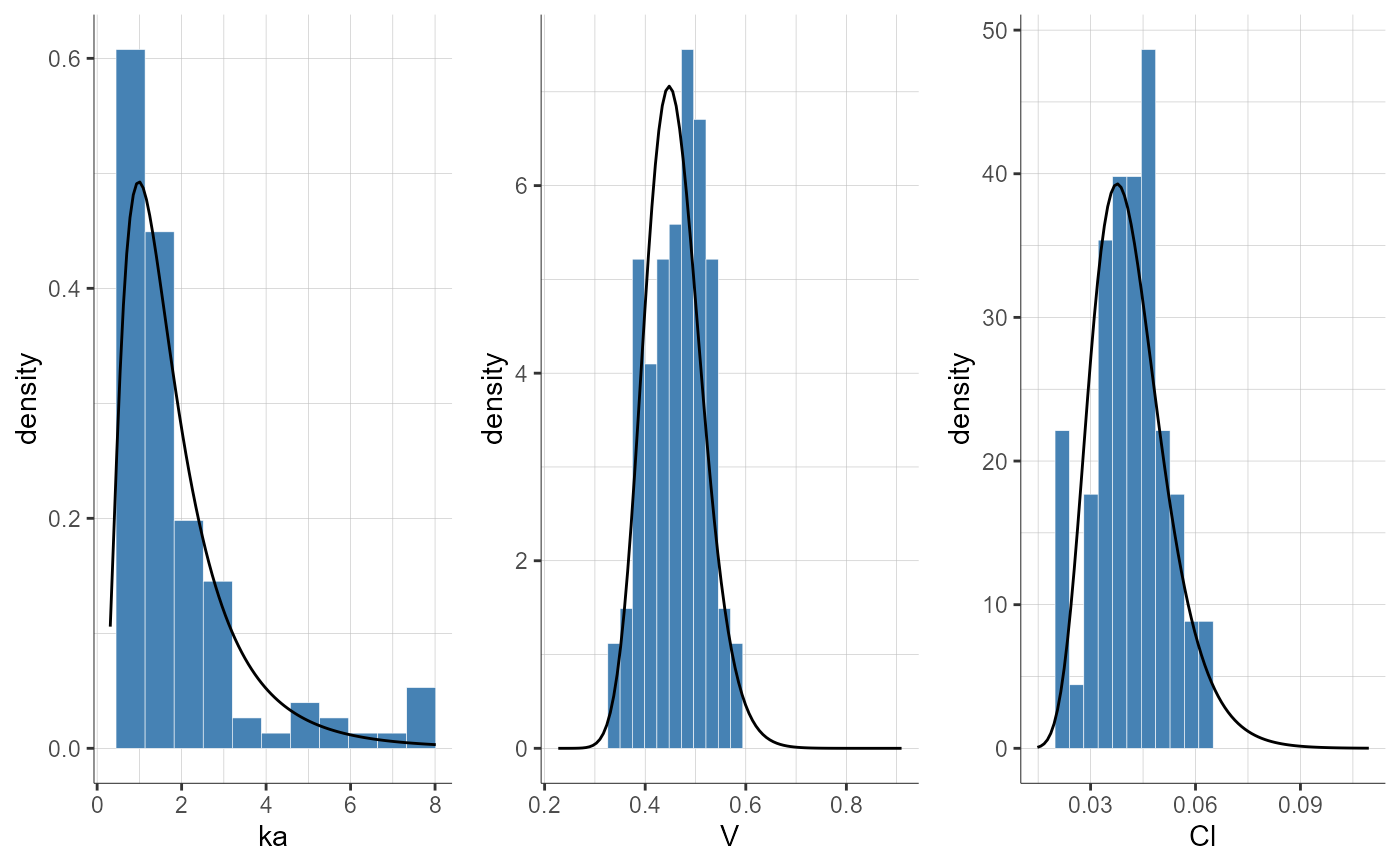
plotParametersDistribution(plot = "pdf")
#> TableGrob (1 x 1) "arrange": 1 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[arrange]
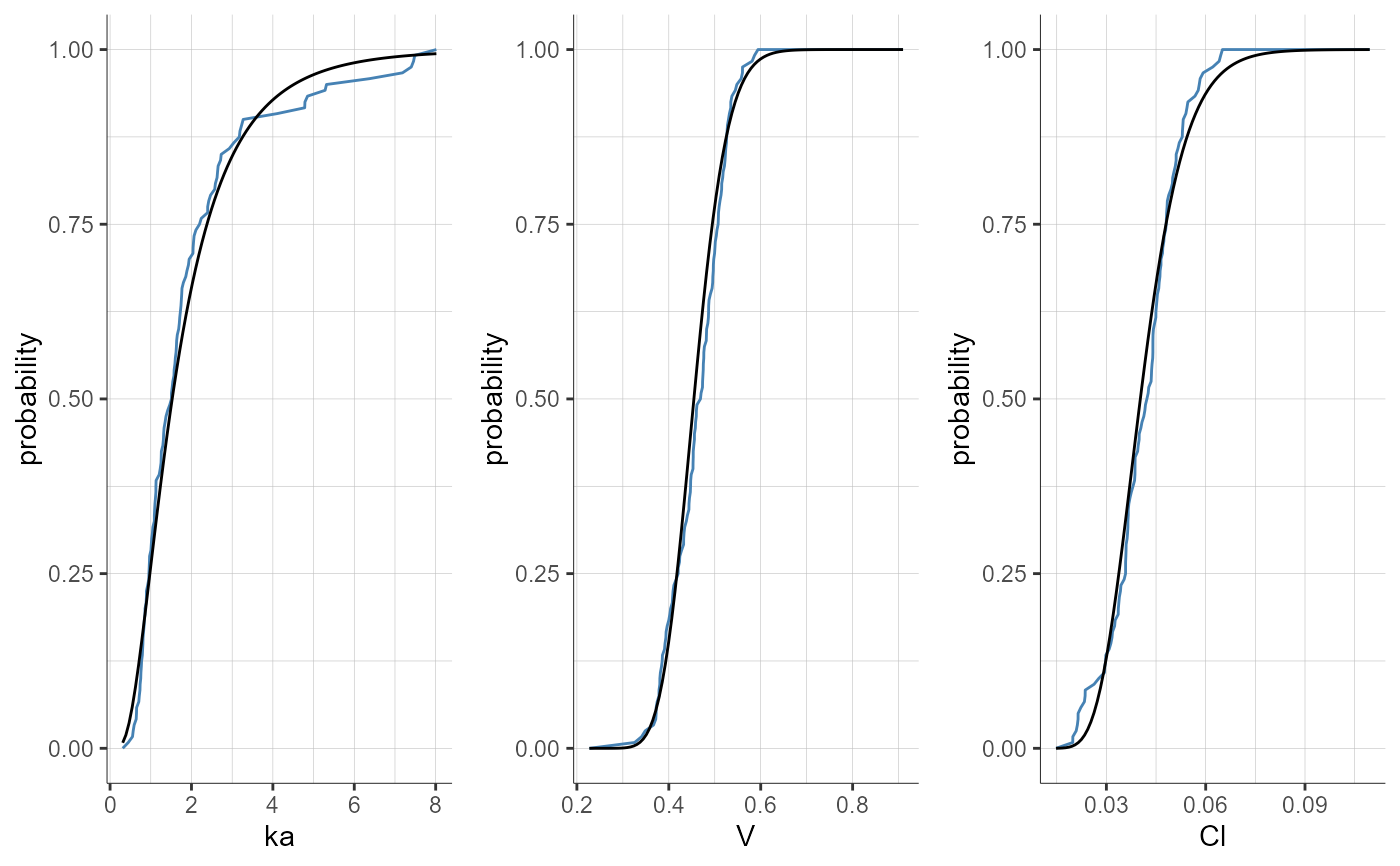
plotParametersDistribution(plot = "cdf")
#> TableGrob (1 x 1) "arrange": 1 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[arrange]
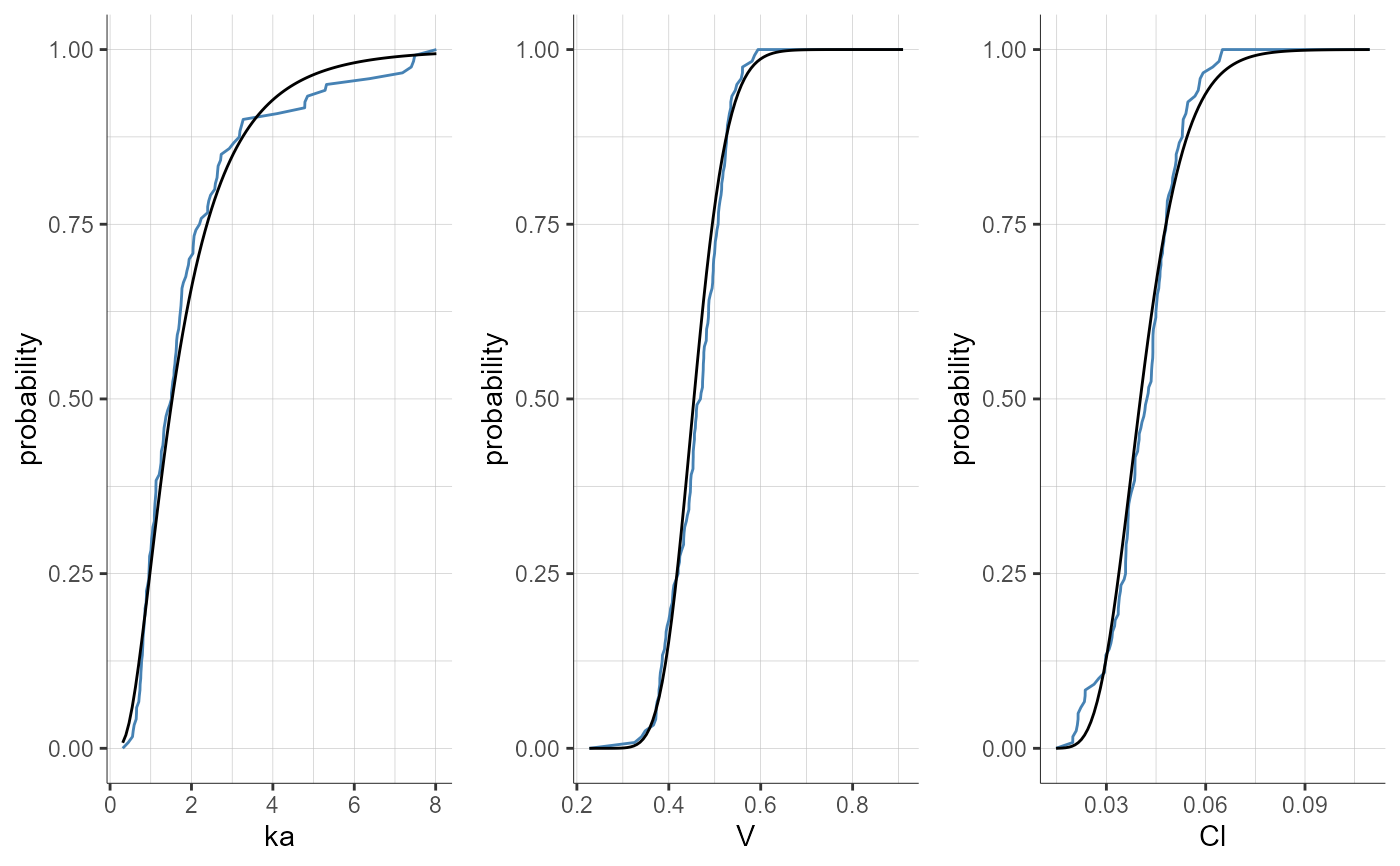
plotParametersDistribution(plot = "cdf", settings = list(indivEstimate = "simulated"))
#> TableGrob (1 x 1) "arrange": 1 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[arrange]
plotParametersDistribution(plot = "pdf", settings = list(theoretical = F))
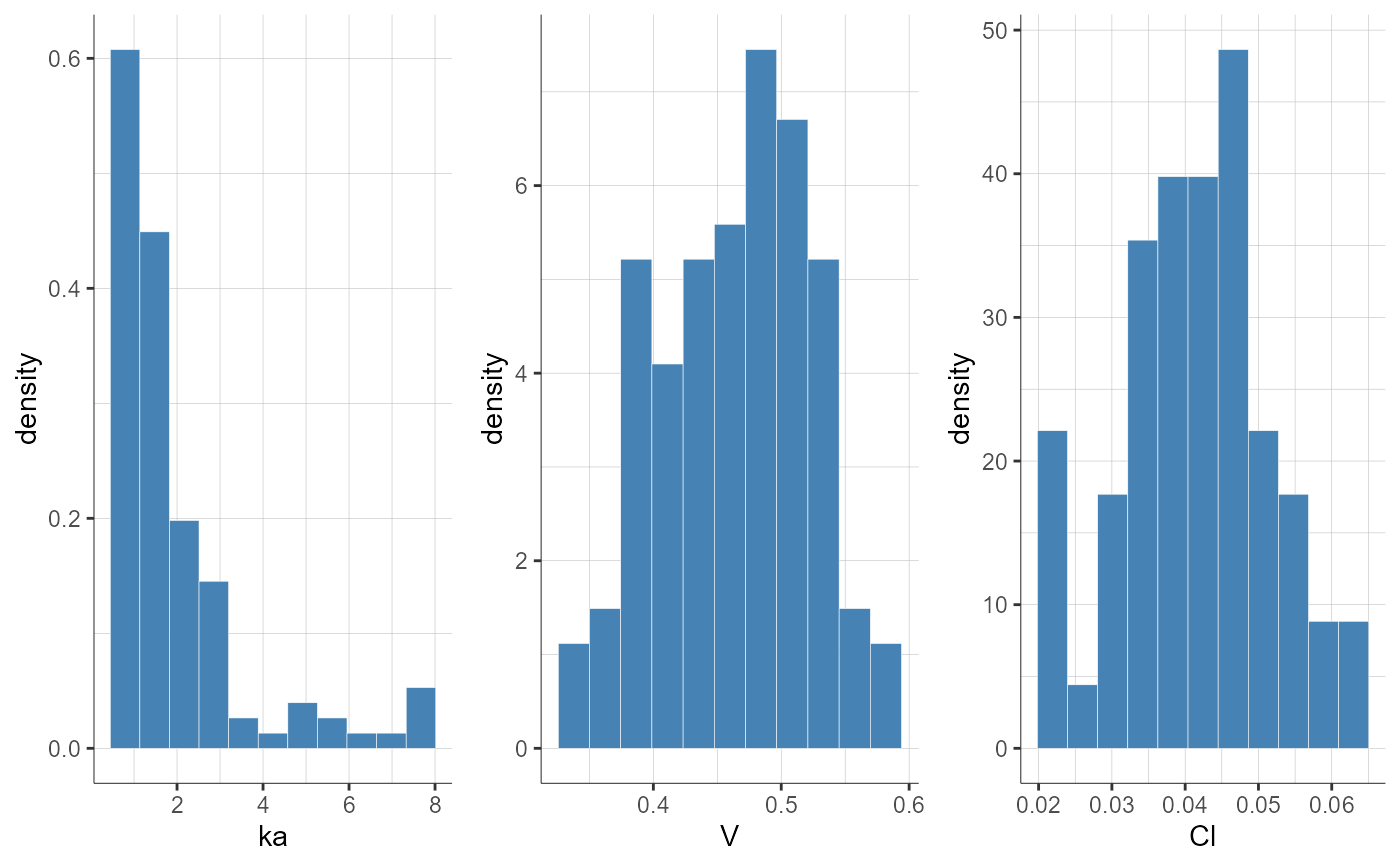
#> TableGrob (1 x 1) "arrange": 1 grobs
#> z cells name grob
#> 1 1 (1-1,1-1) arrange gtable[arrange]

