In the MonolixSuite, the mlxtran language facilitates the description and modeling of time-to-event data. This introduction provides an overview of time-to-event modeling within Monolix, covering various modeling approaches and detailing typical parametric models that are commonly used for this type of analysis.
What is time-to-event data
For time-to-event data, the observations recorded are the times when events occur. This could include the duration from diagnosis to death, or the time between drug administration and the next seizure. In the first scenario, the event is one-time, while in the second it may recur. Events can be:
-
Exactly observed: the event occurs at a specific time
-
Interval censored: the event happens within a known time interval
-
Right censored: the event is unobserved within the study period
Formatting of time-to-event data in the MonolixSuite
The dataset records exactly observed events, interval-censored events, and right-censored events for each individual. Unlike other survival analysis software, MonolixSuite requires specifying the start time of the observation period. This allows for the dataset to be defined using absolute times in addition to durations (if the start time is zero, the records represent durations between the start time and the event).
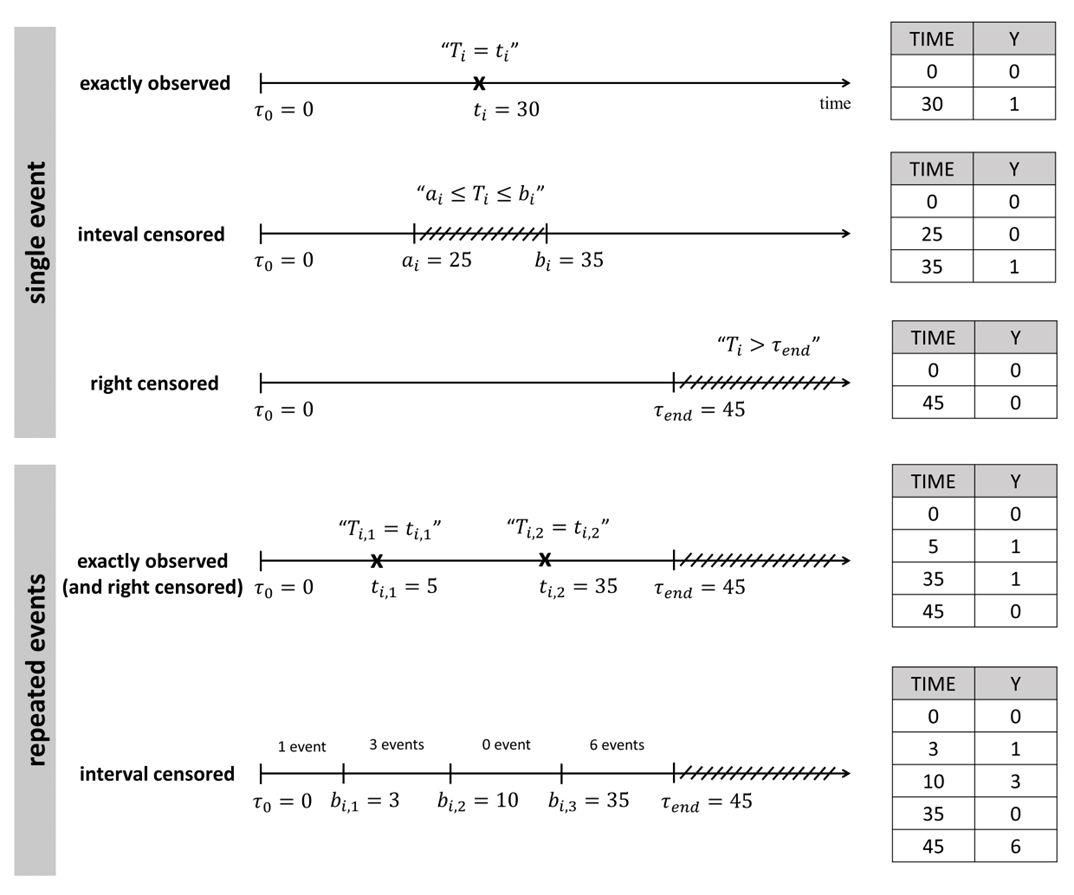

For instance for single events, exactly observed (with or without right censoring), one must indicate the start time of the observation period (Y=0), and the time of event (Y=1) or the time of the end of the observation period if no event has occurred (Y=0). In the following example:
ID TIME Y
1 0 0
1 34 1
2 0 0
2 80 0
The observation period last from starting time t=0 to the final time t=80. For individual 1, the event is observed at t=34, and for individual 2, no event is observed during the period. Thus it is indicated that at the final time (t=80), no event had occurred. Using absolute times instead of durations, we could equivalently write:
ID TIME Y
1 20 0
1 54 1
2 33 0
2 113 0
The durations between start time and event (or end of the observation period) are the same as before, but this time we record the day at which the patients enter the study and the days at which they have events or leave the study. Different patients may enter the study at different times.
Examples for repeated events, and interval censored events are available on the data set documentation page.
Important concepts: hazard and survival
Two functions have a key role in time-to-event analysis: the survival function and the hazard function. The survival function S(t) is the probability that the event happens after time t:
A common way to estimate it non-parametrically is to calculate the Kaplan-Meier estimate. The hazard function
is the instantaneous rate of an event, given that it has not already occurred. Both are linked by the following equation:
The survival funtion
is the exponential function of the antiderivative of the hazard function
Kaplan-Meier estimator
The survival function
is unknown and a typical way to approximate is the non-parametric Kaplan-Meier estimator. It describes the probability that an individual survives until time
, knowing that it survived at any earlier time and, for single events, is given by the following formula:
where
-
– times before t, when at least one event occurred,
-
,
-
– number of individuals at risk, that is who did not experience an event until
The probability that an event occurs
is the ratio of the number of events that have occurred
to the total number of individuals at risk
. The complement of this probability,
, estimates survival. For each time
, the probabilities calculated at all previous times tit_iti, when at least one event occurred, must be multiplied because the total number of individuals at risk has changed. This approach is similar to calculating the probability that a patient survives 2 days, which is the product of the probability of surviving the first day and the conditional probability of surviving the second day given survival of the first day.
Example: A typical time-to-event dataset contains information about the exact times when individuals experienced an event or left the study (drop-out). In this example, five individuals each have two observations: the start time of the observation, which is 0 for all, and the time of the event. If a patient leaves the study, the drop-out time is recorded, but instead of 1 in the observation column, there is 0. This indicates that the individual did not experience an event but survived until the drop-out time. The Kaplan-Meier estimate accounts for situations where not all individuals continue in the study. At each event time, individuals who have dropped out are not counted as at risk and are excluded from the denominator
.
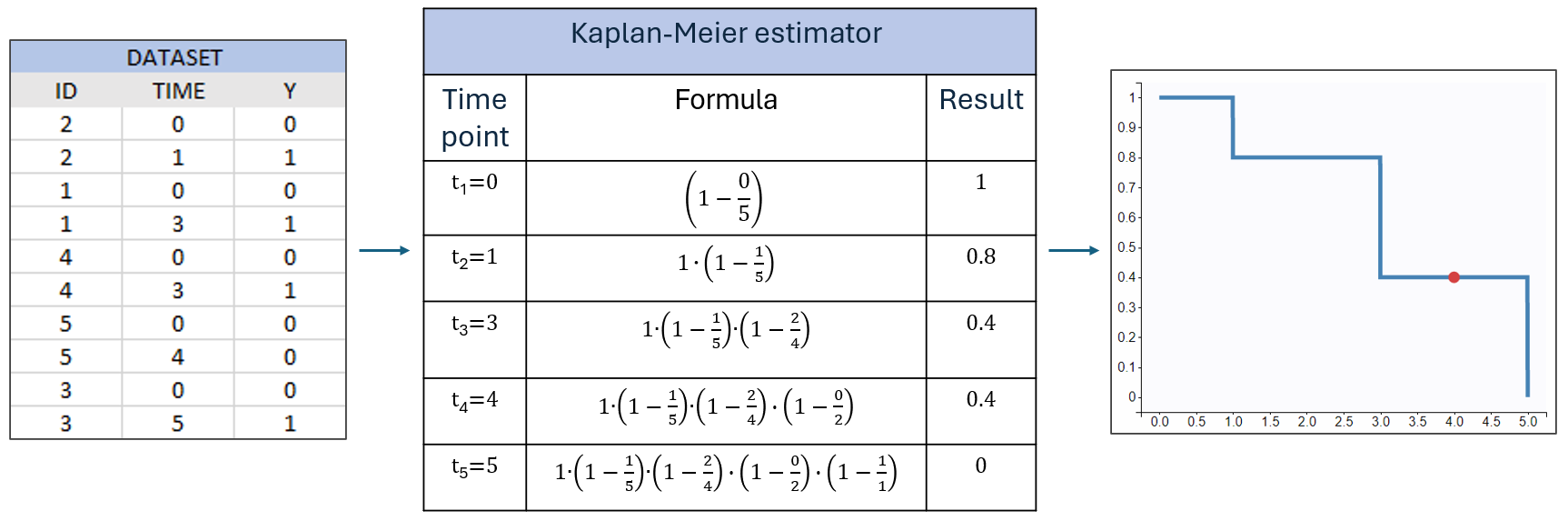
A study starts at time
with no events, so
and the survival curve is 1. Until the next event at
, survival remains constant. At
, one individual experiences an event
, and the total number of individuals at risk
. This results in a 0.2 decrease in survival, reflected as a jump at
in the plot. Survival remains constant until the next event at
, where two events occur. Now,
, having decreased by 1 due to the previous event. The final survival probability is the product of the current probability and all previous probabilities. At
, a drop-out occurs, marked in red, with no event recorded. The survival curve remains constant, and the drop-out is not counted in the risk at the next event at
, where only one individual remains, and one event occurs, resulting in a survival probability of 0.
The Kaplan-Meier estimator correctly handles individuals who leave the study. However, it can be biased when exact event times are unknown and only time intervals are available. For example, if an event is recorded at
, it is unclear whether it occurred exactly at
or earlier. This issue also arises when only the start time and event intervals are provided. In Monolix data visualization, it is assumed that all events are exactly observed, with events occurring at the end of the censored interval if the exact timing is unknown.
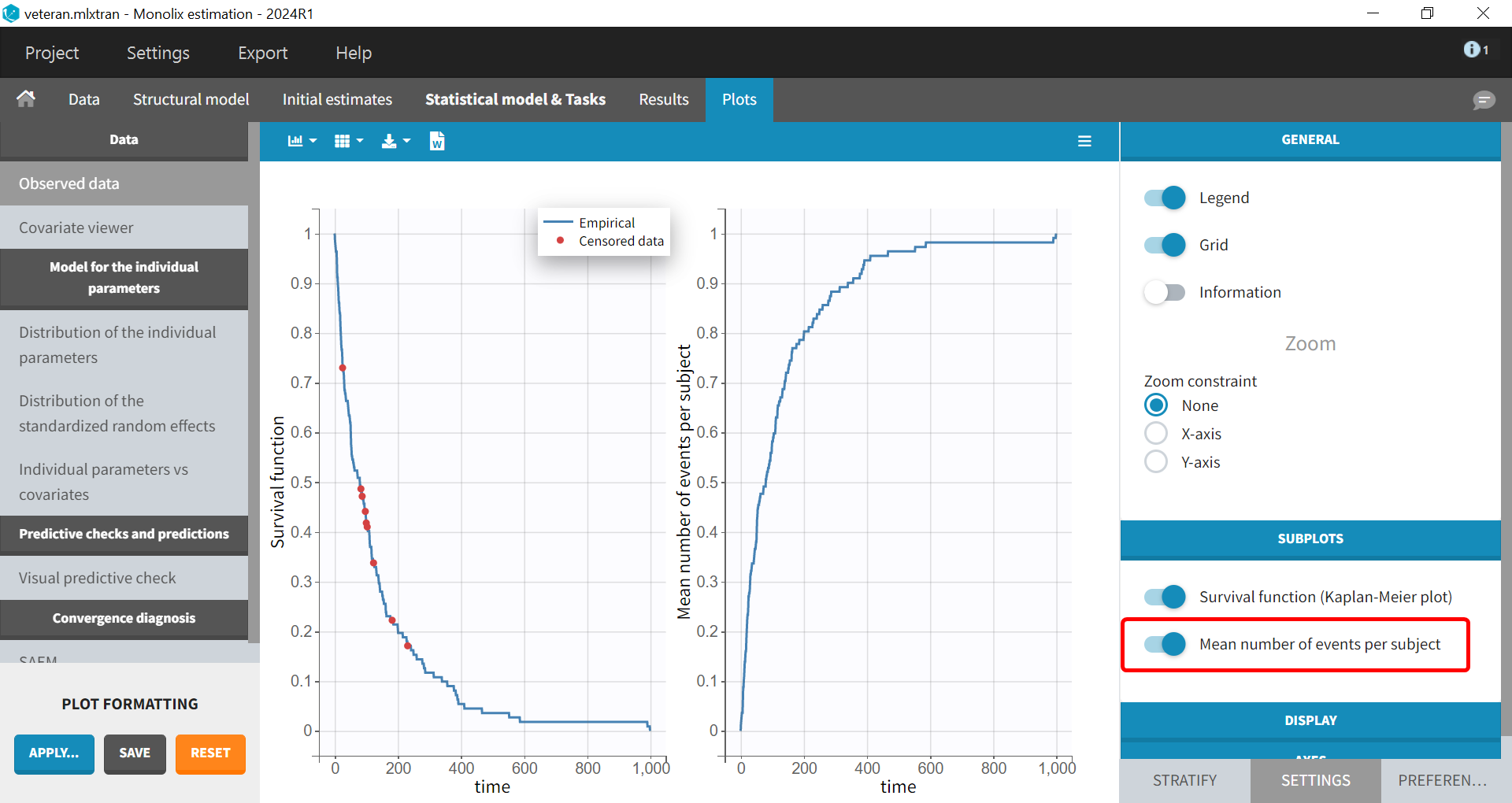
Mean number of events. The Kaplan-Meier estimator can be used also for the analysis of repeated events. The survival curve is estimated for each
-th event separately:
and is used to calculate the mean number of events per individual as a function of time:
It can be visualized in Monolix next to the Survival function by choosing this option from the Subplots settings:
Different types of approaches
Depending on the goal of the time-to-event analysis, different modeling approaches can be used: non-parametric, semi-parametric (Cox models) and parametric.
-
Non-parametric models do not require assumptions on the shape of the hazard or survival. Using the Kaplan-Meier estimate, statistical tests can be performed to check if the survival differs between sub-populations. The main limitations of this approach are that
-
only categorical covariates can be tested and
-
the way the survival is affected by the covariate cannot be assessed.
-
-
Semi-parametric models (Cox models) assume that the hazard can be written as a baseline hazard (that depends only on time), multiplied by a term that depends only on the covariates (and not time). Under this hypothesis of proportional covariate effect, one can analyze the effect of covariates (categorical and continuous) in a parametric way, leaving the baseline hazard undefined.
-
Parametric models require to fully specify the hazard function. If a good model can be found, statistical tests are more powerful than for semi-parametric models. In addition, there is no restrictions on how the covariates affects the hazard. Parametric models can also be easily used for predictions.
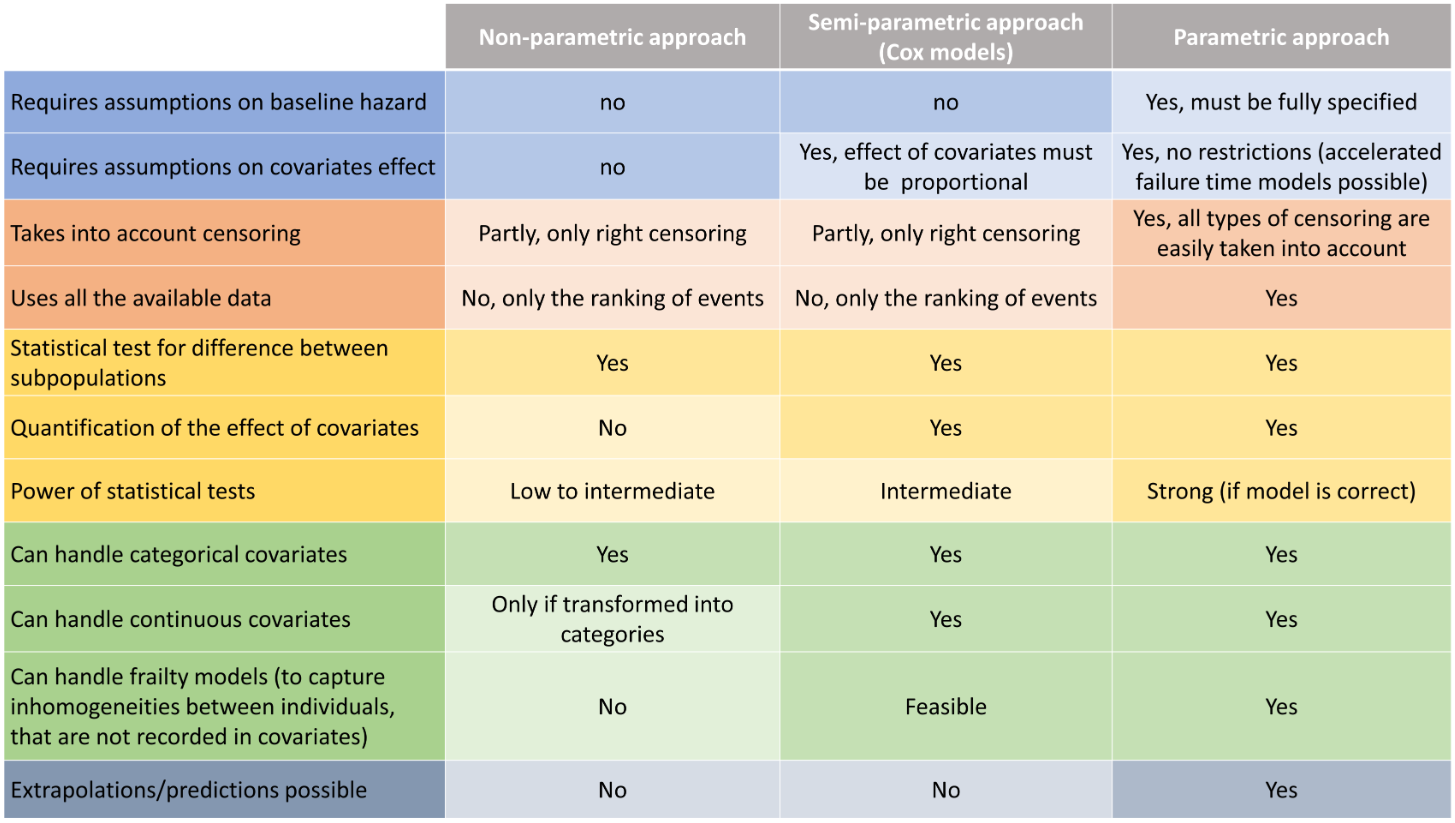
The table below synthesizes the possibilities for the 3 approaches:
Focus on parametric modeling with the MonolixSuite
In the MonolixSuite, the only possible approach is the parametric approach. The model is defined via the hazard function, which in a population approach typically depends on individual parameters:
. With the hazard function, the survival function can easily be computed, as well as the conditional distribution
for various censoring situations (which is required for parameter estimation via SAEM, log-likelihood calculation, etc).
The typical syntax to define the output is the following:
DEFINITION:
Event = {type=event, maxEventNumber=1, hazard=h}
The output Event will be matched to the time-to-event data of the data set. The hazard function h is usually defined via an expression including the input individual parameters. For one-off events, the maximal number of events per individual is 1. It is important to indicate it in the maxEventNumber argument to speed up calculations. To use the model for simulations with Simulx, rightCensoringTime must be given as additional argument. Check here for details.
Note that the hazard can be a function of other variables such as drug concentration or tumor burden for instance (joint PK-TTE or PD-TTE models). An example of the syntax is given here.
Proportional hazard models and hazard ratios
Proportional hazard models are one subclass of models for time-to-event data. In a proportional hazard model, the hazard can be written as the product of a first term (baseline hazard) that depends only on time
and some parameters
, and a second term (link function) that depends only on the covariates
and some parameters
. To ensure a positive hazard, the link is commonly the exponential function.
Hazard ratios for different values of covariates then only depend on
and the covariate, as the baseline hazard cancel out. Thank to this property, proportional hazard models are widely used in the semi-parametric Cox approach. But they can also be implemented using a parametric form in Monolix.
Except for the exponential model (constant hazard), adding covariates on the parameters of the library models (see below for the library) does not lead to a proportional hazard model. Thus, the most convenient approach is to introduce a dummy parameter in the hazard definition, which will carry the covariates.
For instance, the log-logistic model from the library can easily be modified to add the dummy parameter h_cov, both in the input list and in the hazard definition:
[LONGITUDINAL]
input = {Te, s, h_cov}
EQUATION:
h = s/Te * (t/Te)^(s-1) / (1+(t/Te)^s) * h_cov
DEFINITION:
Event = {type=event, maxEventNumber=1, hazard=h}
OUTPUT:
output = {Event}
The statistical model must then be defined in the following way:
-
remove the random effects on
h_cov. Keep the default log-normal distribution. -
fix the value of the fixed effect
h_cov_popto 1 -
add untransformed covariates on
h_cov.
Using a log-normal distribution and untransformed covariates will lead to the following hazard. For the example, we assume that AGE and TRT have been added on h_cov:
The estimated betas and their standard errors are displayed in the Monolix results. Note that the Wald test is also given and that automatic covariate search strategies can be applied.
This strategy with the dummy parameter does not allow to consider interaction between covariates. If this is needed, the covariates can be passed as regressor and the covariate model can be implemented directly in the structural model.
Library of parametric models for time-to-event data
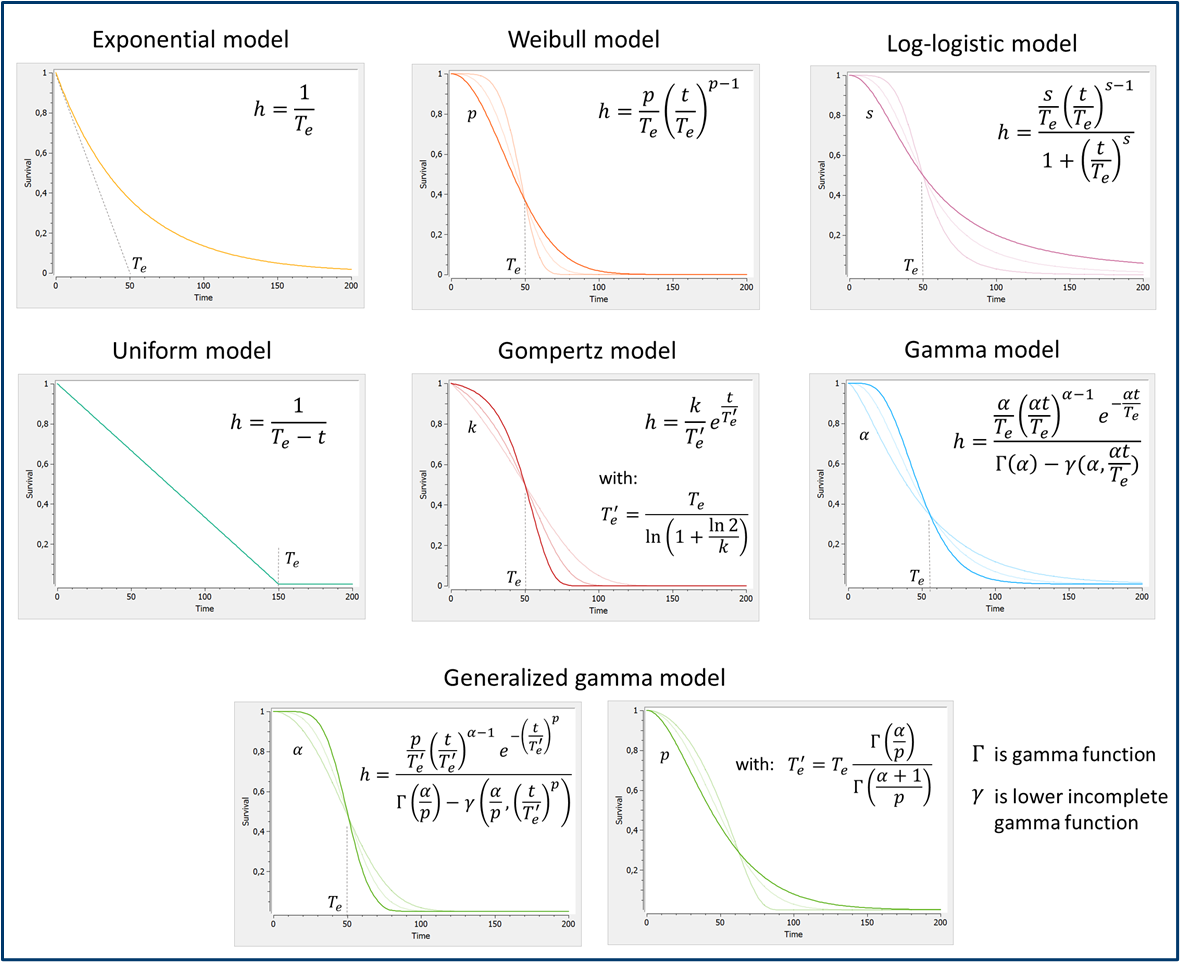
To describe the various shapes that the survival Kaplan-Meier estimate can take, several hazard functions have been proposed. Below we display the survival curves, for the most typical hazard functions:
A few comments:
-
We have reparametrized
as a function of
to better separate the effects of the scale parameter (characteristic time) and the shape parameter (shape of the curve).
-
All parameters are positive. If we assume inter-individual variability, a log-normal distribution is usually appropriate.
The table below summarizes the number of parameters and typical parameter values:
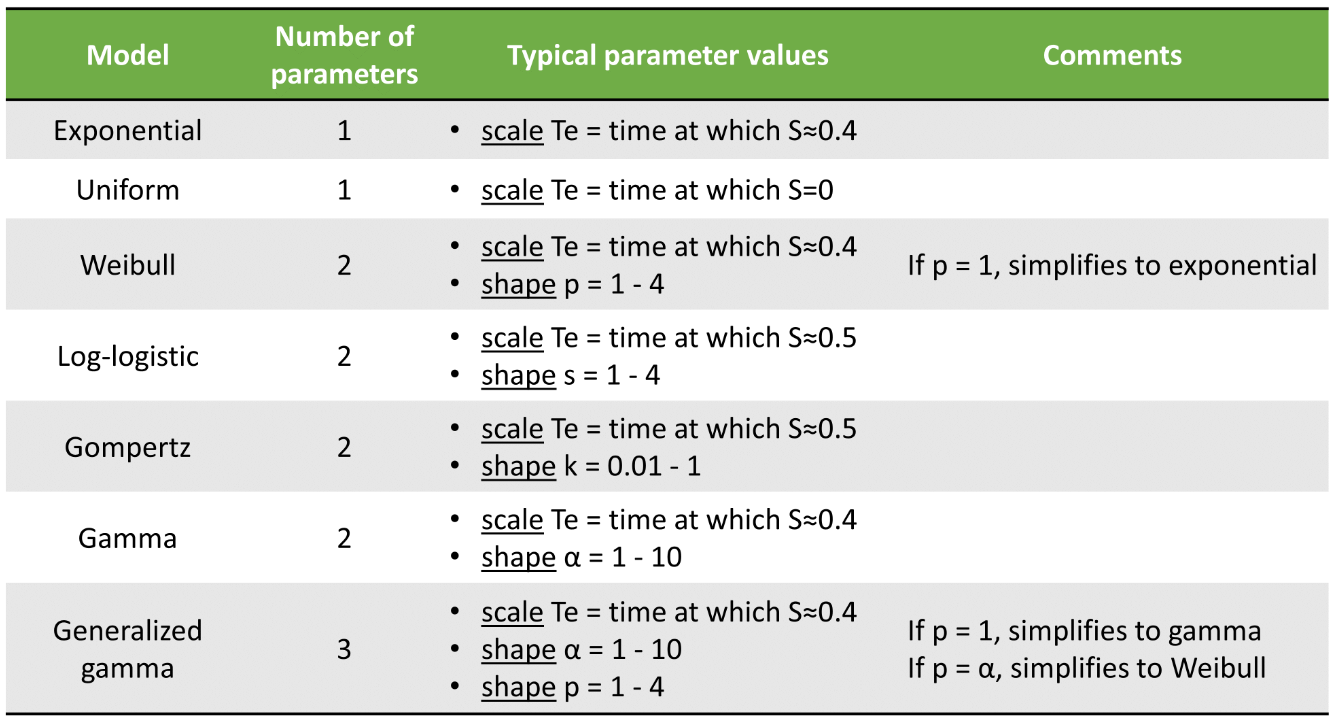
For each model, we can in addition consider a delay del as additional parameter. The delay will simply shift the survival curve to the right (later times). For
, the survival is
.
Downloads:
All models are available as Mlxtran model file in the TTE library. Each model can be with/without delay and for single/repeated events. For performance reasons, it is important to choose the file ending with ‘_singleEvent.txt’ if you want to model one-off events (death, drop-out, etc).
Case studies
Two case studies show the modeling and simulation workflow for TTE data:

