[PKanalix] Generate NCA data plots
Plot the observed data as used for the NCA calculations
Usage
plotNCAData(settings = list(), stratify = list(), preferences = list())
Arguments
settings List with the following settings:dots (logical) - If TRUE, individual observations are displayed as dots (default TRUE). lines (logical) - If TRUE, individual observations are connected by lines (default FALSE if NCA has been run using sparse data option, TRUE either). mean (logical) - If TRUE, mean (geometric or arithmetic) of observations is displayed (default TRUE). error (logical) If TRUE, error bar presenting standard deviation or standard error are displayed (default TRUE). meanMethod (character) - When mean is set to TRUE, display arithmetic mean ("arithmetic", default) or geometric mean ("geometric"). Forced to 'arithmetic' if NCA has been run using sparse data option. errorMethod (character) - When error is set to TRUE, display standard deviation ("standardDeviation", default) or standard error ("standardError"). Forced to 'standardDeviation' if NCA has been run using sparse data option. useCensored (logical) If TRUE, censored observations are used to compute mean and error (default TRUE). Not available if NCA has been run using sparse data option. binLimits (logical) - If TRUE, bins limits are displayed as vertical lines (default FALSE). Not available if NCA has been run using sparse data option. binsSettings a list of settings for time axis binning for observation statistics computation. Not available if NCA has been run using sparse data option.criteria (character) - Binning criteria, one of 'equalwidth', 'equalsize', or 'leastsquare' (default) methods. is.fixedNbBins (logical) - If TRUE, a fixed number of bins is used (see nbBins). If FALSE (default), the number of bins is optimized using binRange and nbBinData. nbBins (integer) - Number of bins (default 10). Ignored if is.fixedNbBins=FALSE. binRange (vector(integer, integer)) - Minimum and maximum number of bins when bins are optimized (default c(5, 100)). nbBinData (vector(integer, integer)) - Minimum and maximum number of data points per bin when bins are optimized (default and c(3, 200)). cens (logical) - If TRUE, censored observations are displayed as dots (default TRUE). dosingTimes (logical) - If TRUE, dosing times are displayed as vertical lines (default FALSE). legend (logical) - If TRUE, plot legend is displayed (default FALSE). grid (logical) - If TRUE, plot grid is displayed (default TRUE). xlog (logical) - If TRUE, log-scale on x axis is used (default FALSE). ylog (logical) - If TRUE, log-scale on y axis is used (default FALSE). xlab (character) - Label on x axis (default "time"). ylab (character) - Label on y axis (default obsName). ncol (integer) - Number of columns when using split (default 4). xlim (c(double, double)) - Limits of the x axis. ylim (c(double, double)) - Limits of the y axis. fontsize (integer) - Font size of text elements (default 11). units (logical) - If TRUE, units are added in axis labels (default TRUE). scales (character) Should scales be fixed ("fixed"), free ("free", default), or free in one dimension ("free_x", "free_y"). stratify List with the stratification arguments:groups - Definition of stratification groups. By default, stratification groups are already defined as one group for each category for categorical covariates, and two groups of equal number of individuals for continuous covariates. To redefine groups, for each covariate to redefine, specify a list with:namecharactercovariate name (e.g "AGE")definition(vector(continuous) || list>(categorical))For continuous covariates, vector of break values (e.g c(35, 65)). For categorical covariates, groups of categories as a list of vectors(e.g list(c("study101"), c("study201","study202"))) split (vector) - Vector of covariates used to split (i.e facet) the plot (by default no split is applied). For instance c("FORM","AGE"). mergedSplits (logical) - When "split" is used and mean=T, should the means of the different groups be displayed on the same plot (TRUE) or on different subplots (FALSE, default). Not available for count/categorical data. When mergedSplits=T, the "color" argument is ignored and the coloring is done according to the splitted groups. filter (list< list> >) - List of pairs containing a covariate name and the vector of indexes or categories (for categorical covariates) of the groups to keep (by default no filtering is applied). For instance, list("AGE",c(1,3)) to keep the individuals belonging to the first and third age group, according to the definition in groups. For instance, list("FORM","ref") using the category name for categorical covariates. color (vector) - Vector of covariates used for coloring or "id" (by default no coloring is applied). For instance c("FORM","AGE") or "id". colors - Vector of colors to use when color argument is used. Takes precedence over colors defined in preferences. For instance c("#ebecf0","#cdced1","#97989c"). individualSelection - Ids to display (by default the 12 first ids are displayed) defined as:indices (vector) - Indices of the individuals to display (by default, the 12 first individuals are selected). If occasions are present, all occasions of the selected individuals will be displayed. Takes precedence over ids. For instance c(5,6,10,11). isRange (logical) - If TRUE, all individuals whose index is inside [min(indices), max[indices]] are selected (FALSE by default). Forced to FALSE if ids is defined. ids (vector) - Names of the individuals to display. If occasions are present, all occasions of the selected individuals will be displayed. For instance c("101-01","101-02","101-03"). If ids are integers, can also be c(1,3,6). Ignored if indices is defined. preferences (optional) preferences for plot display, run getPlotPreferences("plotNCAData") to check available options.
Value
A ggplot object
See also
Examples
initializeLixoftConnectors(software = "pkanalix")
project <- paste0(getDemoPath(), "/2.case_studies/project_aPCSK9_SAD.pkx")
loadProject(project)
setNCASettings(blqMethodBeforeTmax="missing",blqMethodAfterTmax="LOQ")
runNCAEstimation()
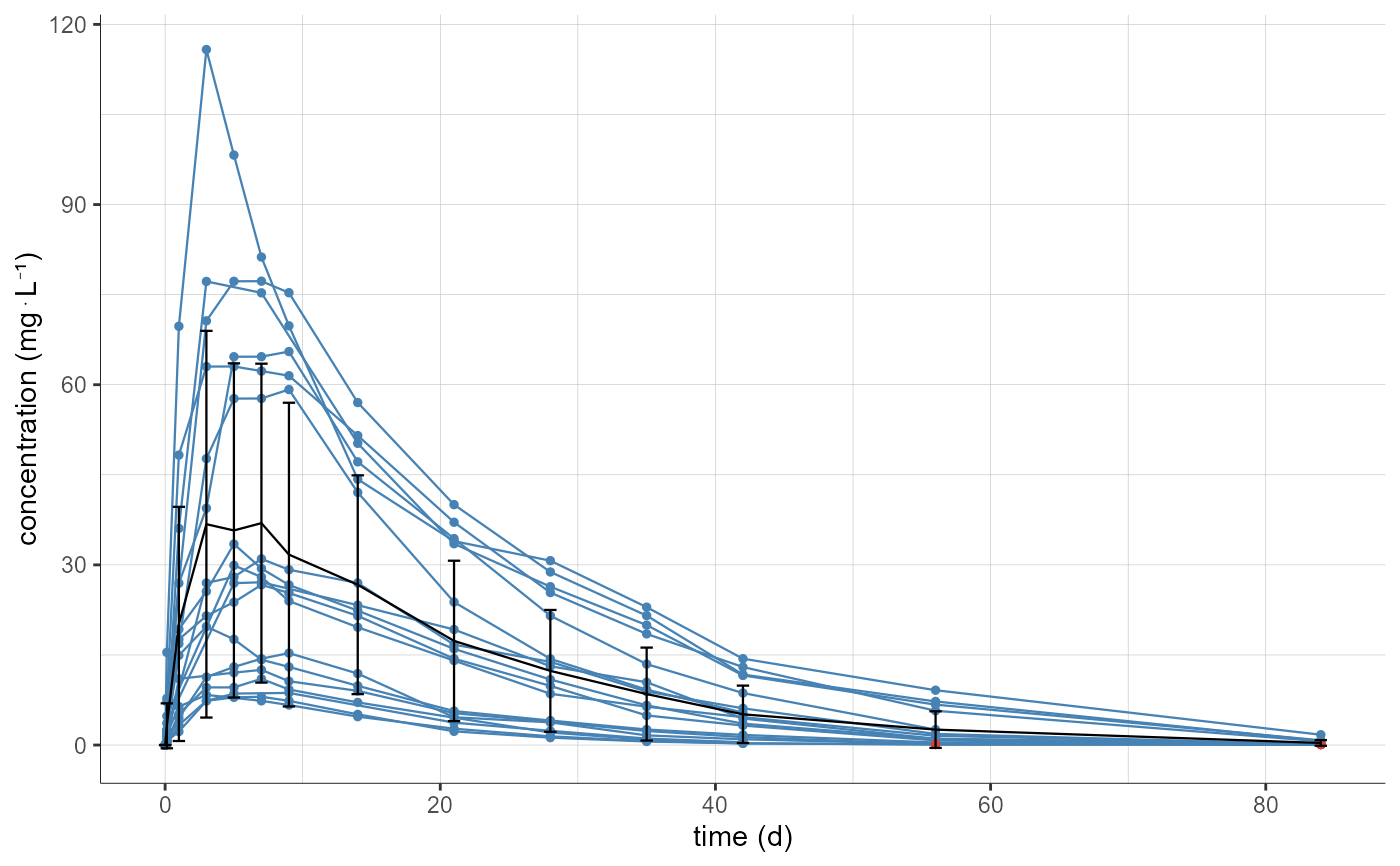
# by default, individual profiles and mean curve are displayed
plotNCAData()
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
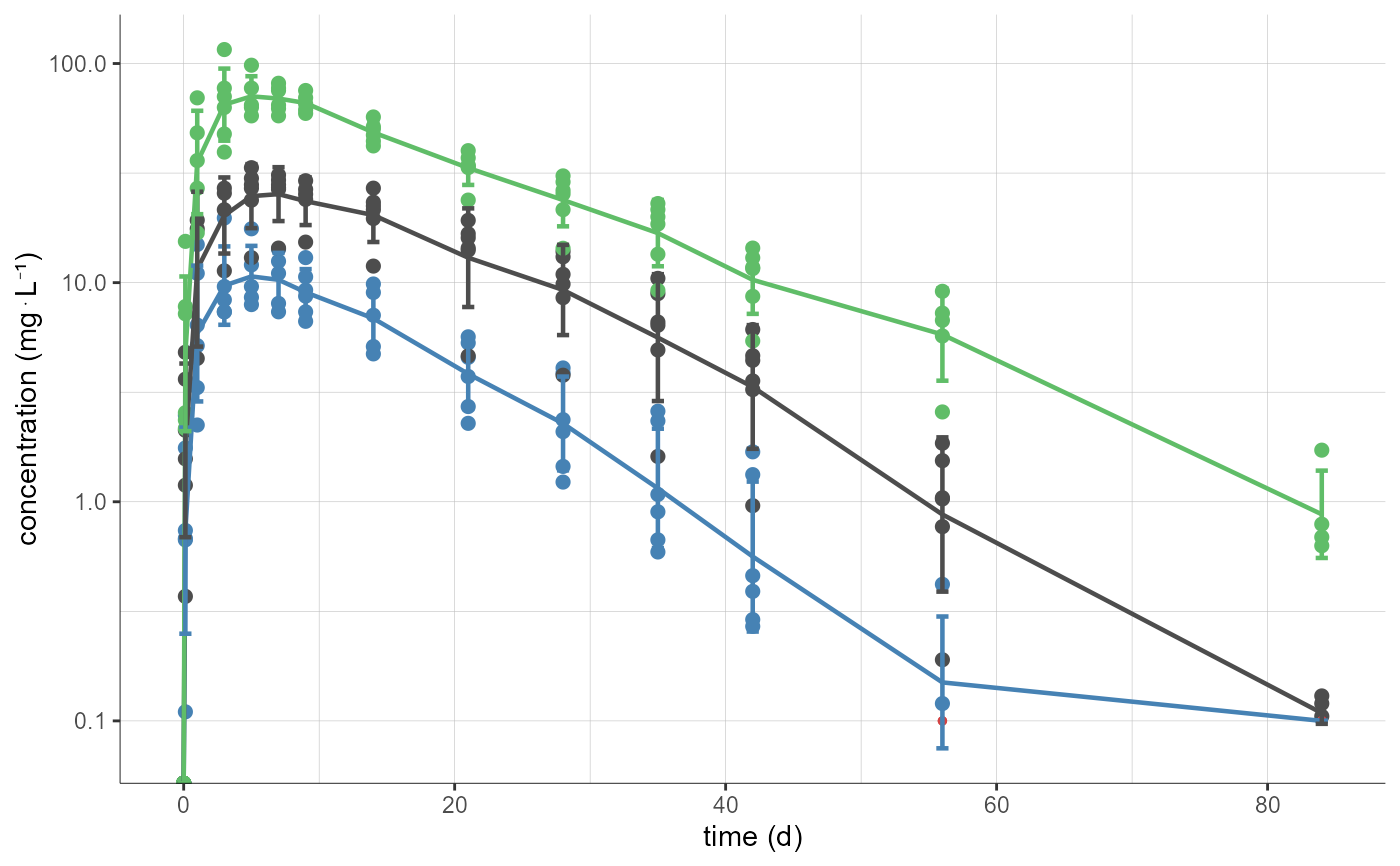
# displaying dots and mean curve by dose group, merged on a single plot
plotNCAData(settings=list(lines=F,
mean=T,
meanMethod="geometric",
ylog=T),
stratify=list(split="DOSE_mg",
mergedSplits=T),
preferences=list(observationStatistics=list(lineWidth=0.8),
obs=list(radius=2)))
#> Warning: log-10 transformation introduced infinite values.
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
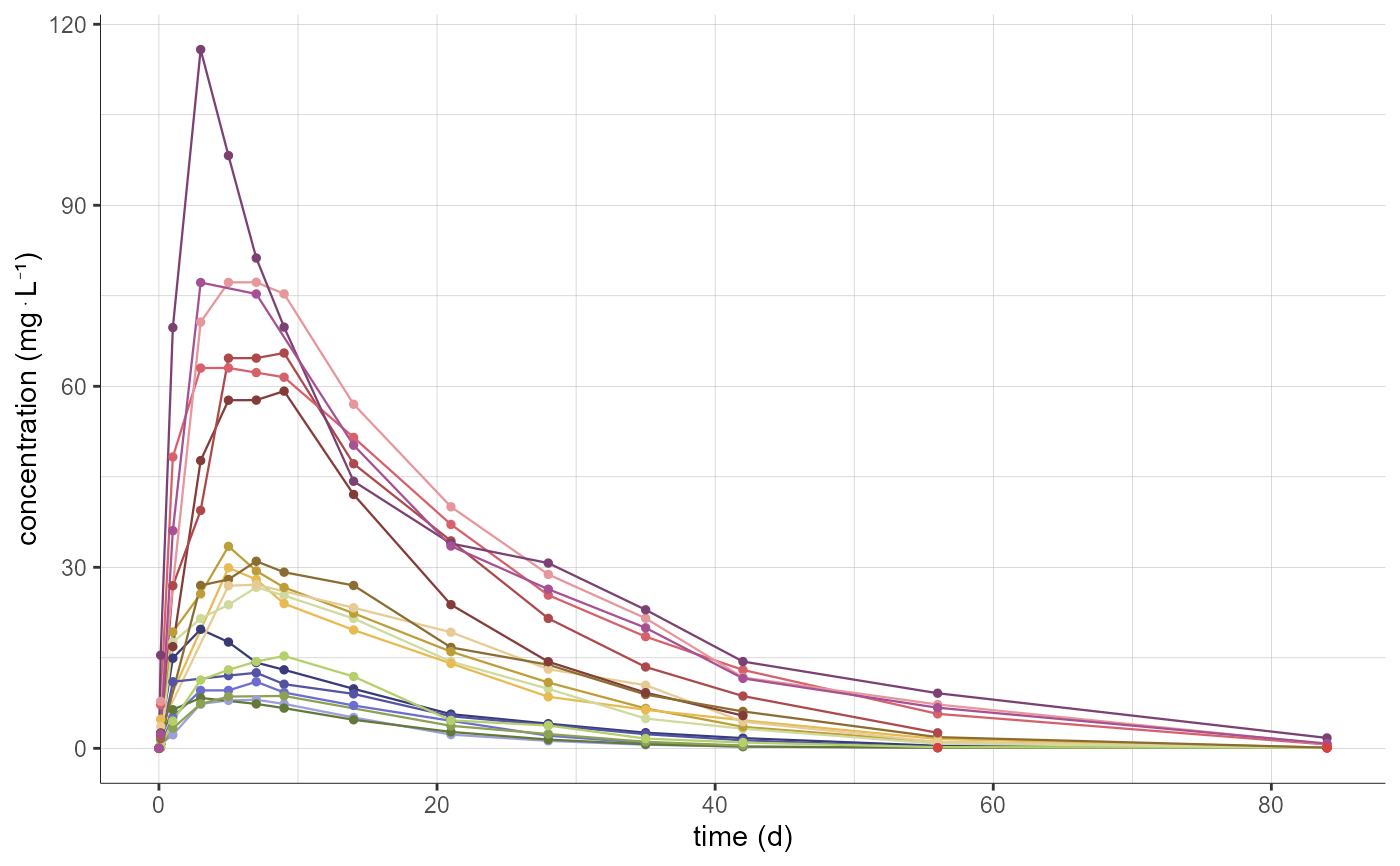
# coloring by ID, without mean curve
plotNCAData(settings=list(mean=F,error=F),
stratify = list(color=c("id")))
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
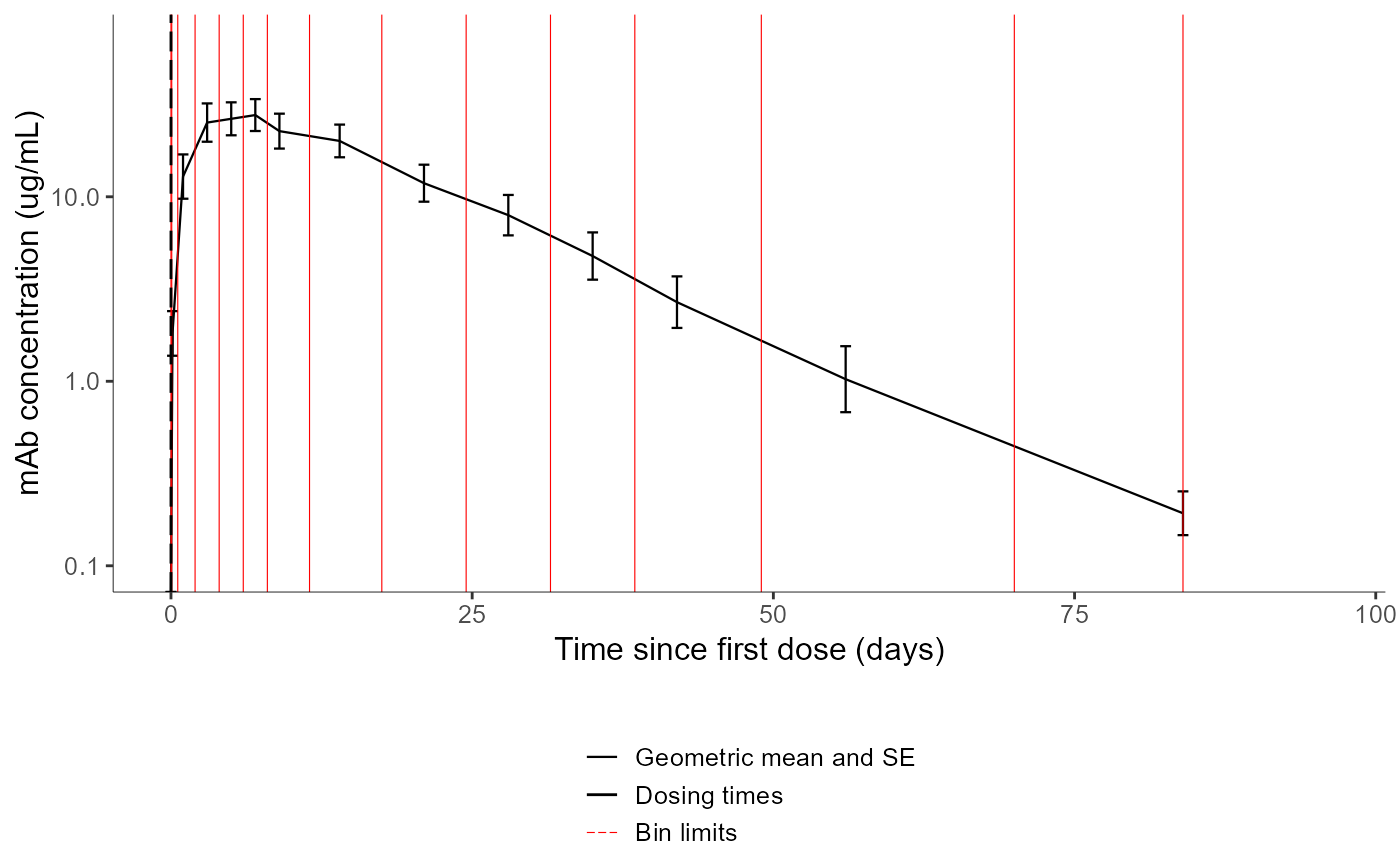
# changing the settings to display only the mean curve with SE, with bin limits and dosing times
plotNCAData(settings=list(dots=F,
lines=F,
mean=T,
error=T,
meanMethod="geometric",
errorMethod="standardError",
useCensored=T,
binLimits=T,
binsSettings=list(criteria="leastsquare", is.fixedNbBins=T, nbBins=20),
cens=F,
dosingTimes=T,
legend=T,
grid=F,
xlog=F,ylog=T,
xlab="Time since first dose (days)",
ylab="mAb concentration (ug/mL)",
xlim=c(0,96),
ylim=c(0.1,70),
fontsize=12,
units=F))
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
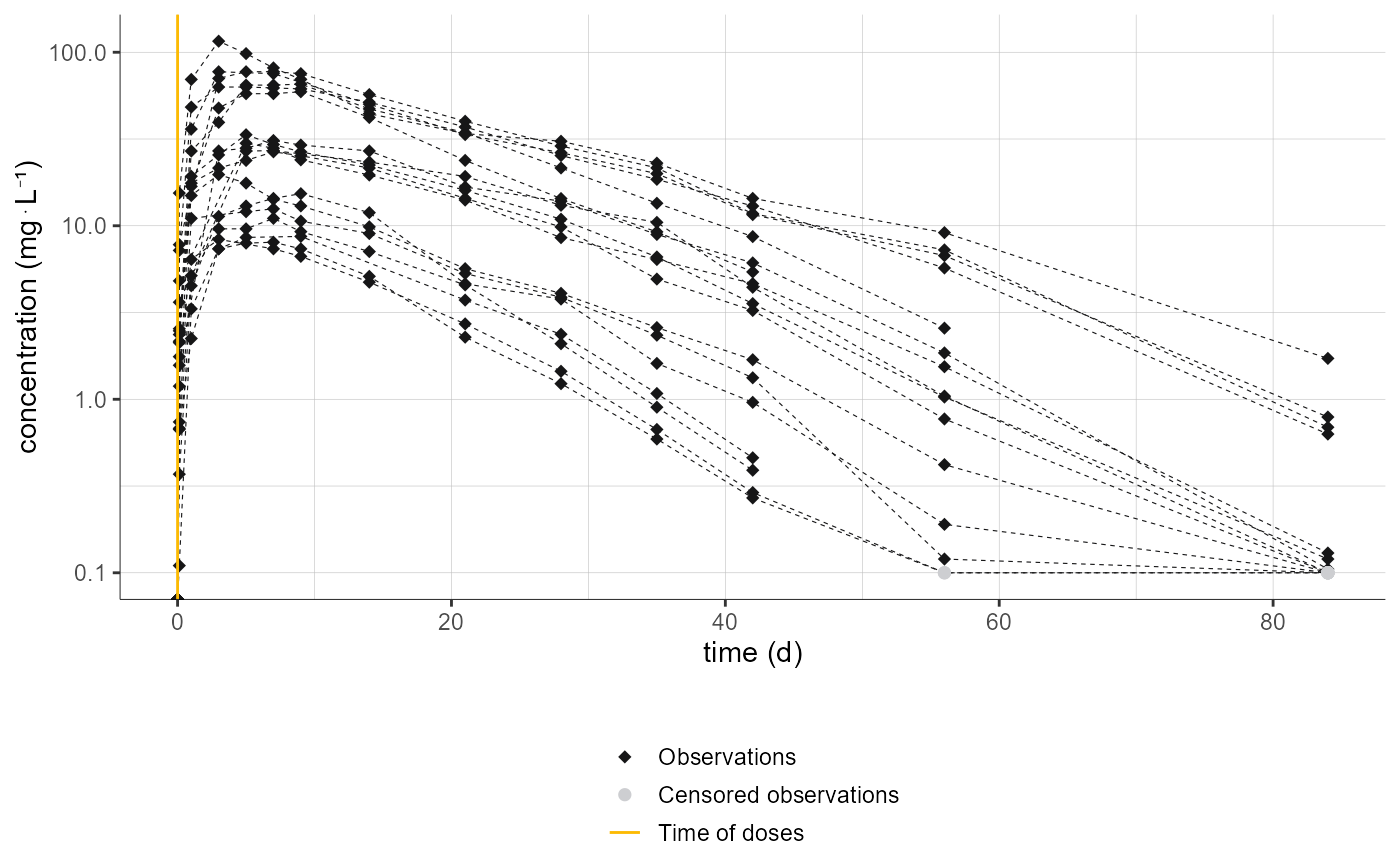
# changing preferences for observations, censored observations and bin limits
plotNCAData(settings=list(dots=T, lines=T, legend=T, dosingTimes=T, mean=F, error=F, ylog=T, cens=T),
preferences=list(obs=list(color="#161617",
radius=2,
shape=18,
lineWidth=0.2,
lineType="dashed",
legend="Observations"),
censObs=list(color="#cdced1",
radius=2,
shape=16,
legend="Censored observations"),
dosingTimes=list(color="#fcba03",
lineWidth=0.5,
lineType="solid",
legend="Time of doses")))
#> Warning: log-10 transformation introduced infinite values.
#> Warning: log-10 transformation introduced infinite values.
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
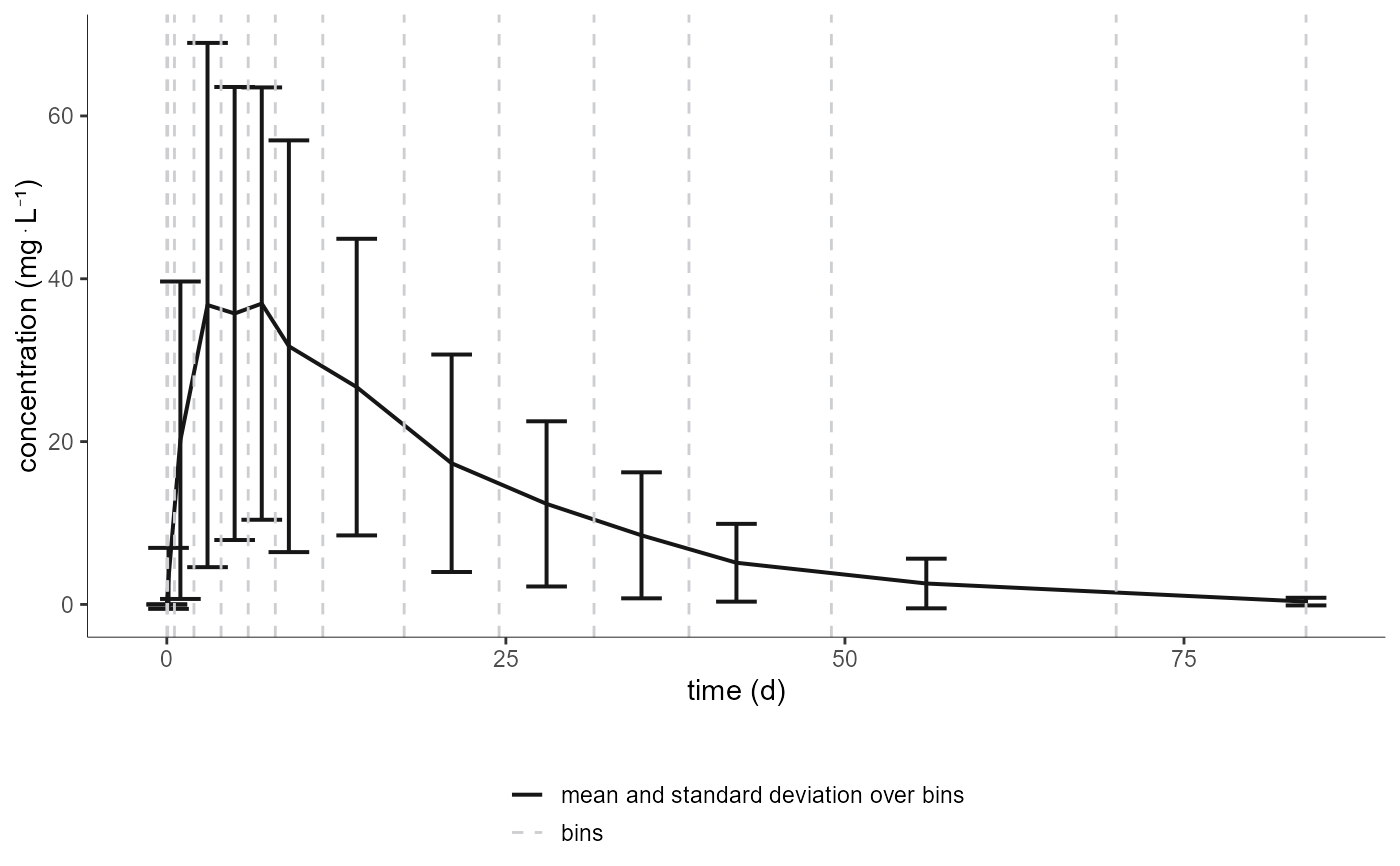
# changing preferences for mean and bin limits
plotNCAData(settings=list(dots=F, lines=F, legend=T, binLimits=T, grid=F),
preferences=list(observationStatistics=list(color="#161617",
whiskersWidth=3,
lineWidth=0.7,
lineType="solid",
legend="mean and standard deviation over bins"),
binsValues=list(color="#cdced1",
lineWidth=0.5,
lineType="dashed",
legend="bins")))
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
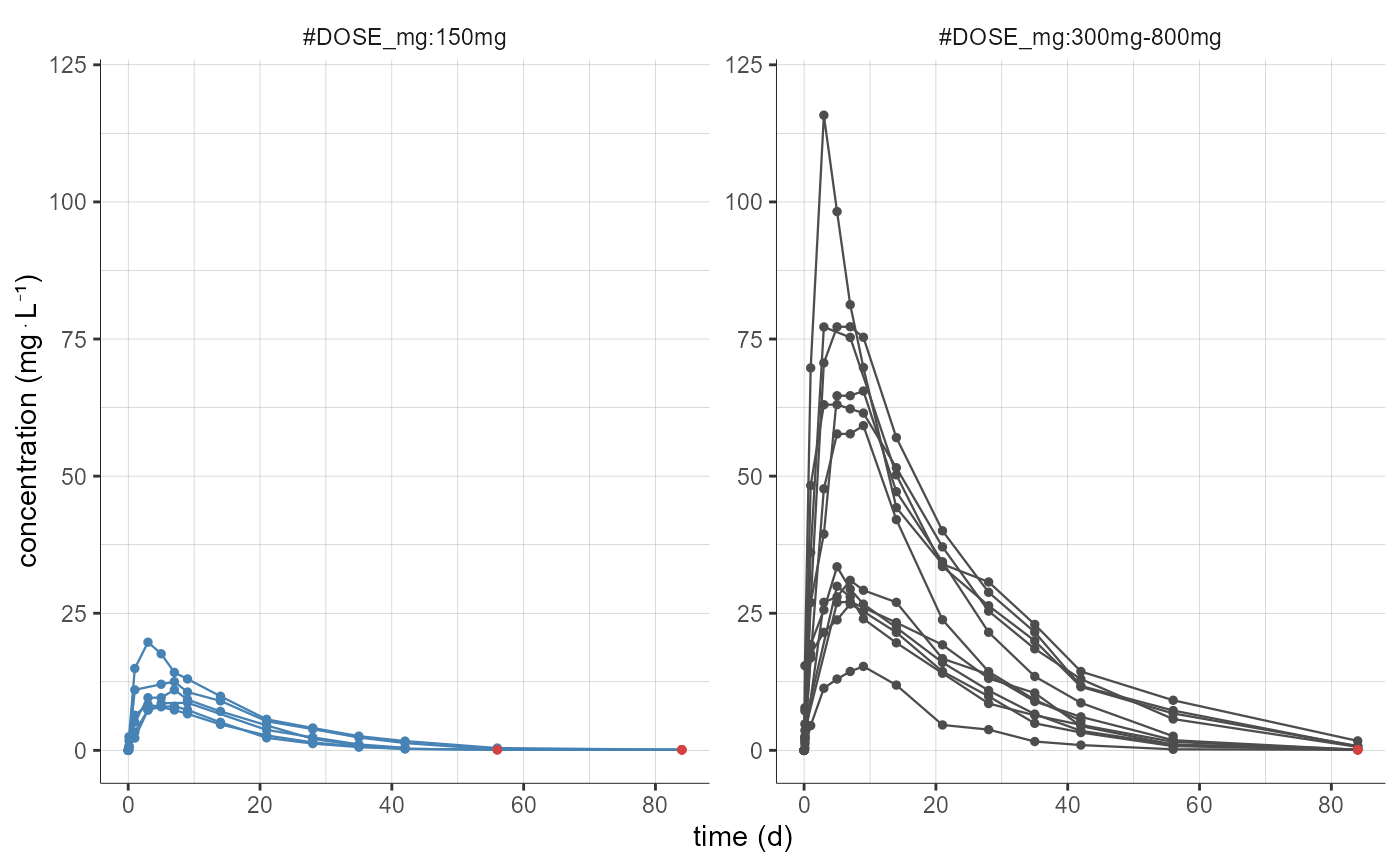
# color and split by DOSE_mg but grouping two doses levels together
plotNCAData(settings=list(mean=F,error=F,ylim=c(0,120)),
stratify = list(groups=list(name="DOSE_mg",definition=list(c("150mg"), c("300mg","800mg"))),
color="DOSE_mg",
split="DOSE_mg"))
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
#============= projects with several covariates
project <- file.path(getDemoPath(), "/2.case_studies/project_Theo_extravasc_SD.pkx")
loadProject(project)
runNCAEstimation()
# defining groups for AGE and HT, coloring by HT and filtering by AGE
plotNCAData(settings=list(mean=F,error=F),
stratify = list(groups=list(list(name="AGE", definition=c(24, 34)),
list(name="HT", definition=c(184.5))),
color="HT",
colors=c("#cdced1","#161617"),
filter=list("AGE",c(1,3))))
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
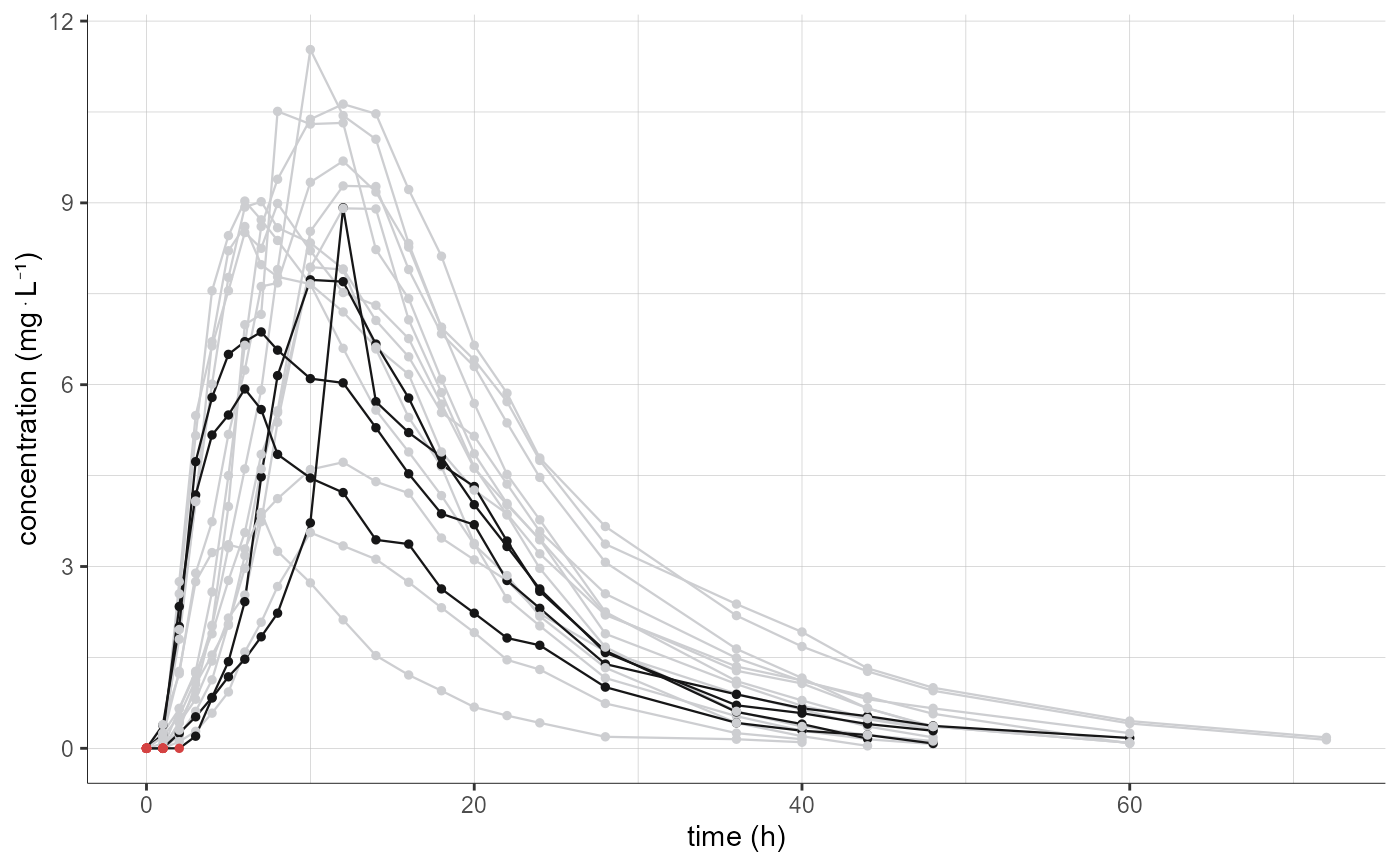
# filter to keep only second sequence (TR) and FORM=test
plotNCAData(settings=list(mean=F,error=F),
stratify = list(filter=list(list("SEQ",2),list("FORM","test"))))
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
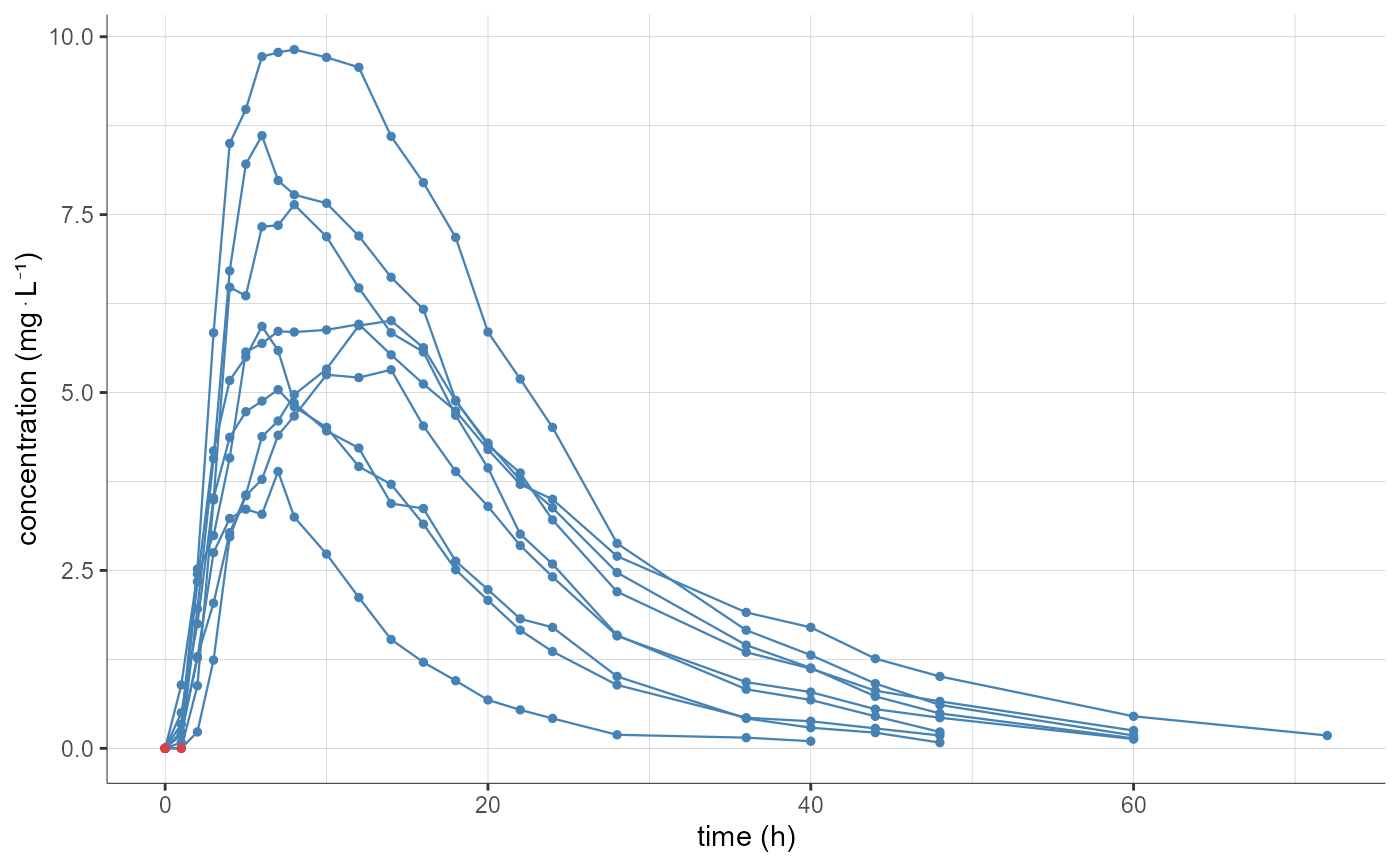
#============= project with sparse calculations
project <- paste0(getDemoPath(), "/1.basic_examples/project_sparse.pkx")
loadProject(project)
runNCAEstimation()
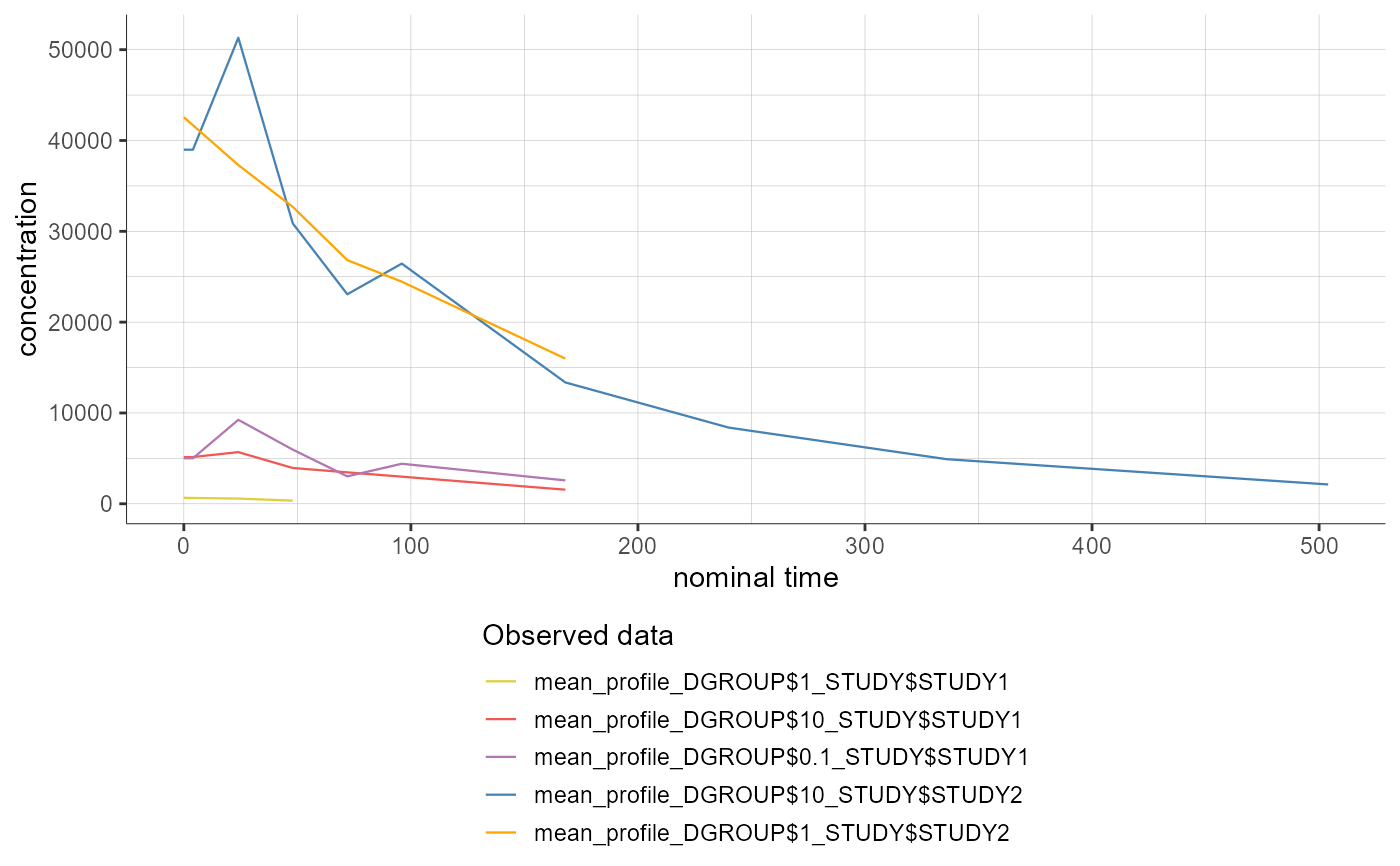
# mean profiles are already calculated for each stratification group used for the sparse calculations
# stratify argument cannot be used
plotNCAData()
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.
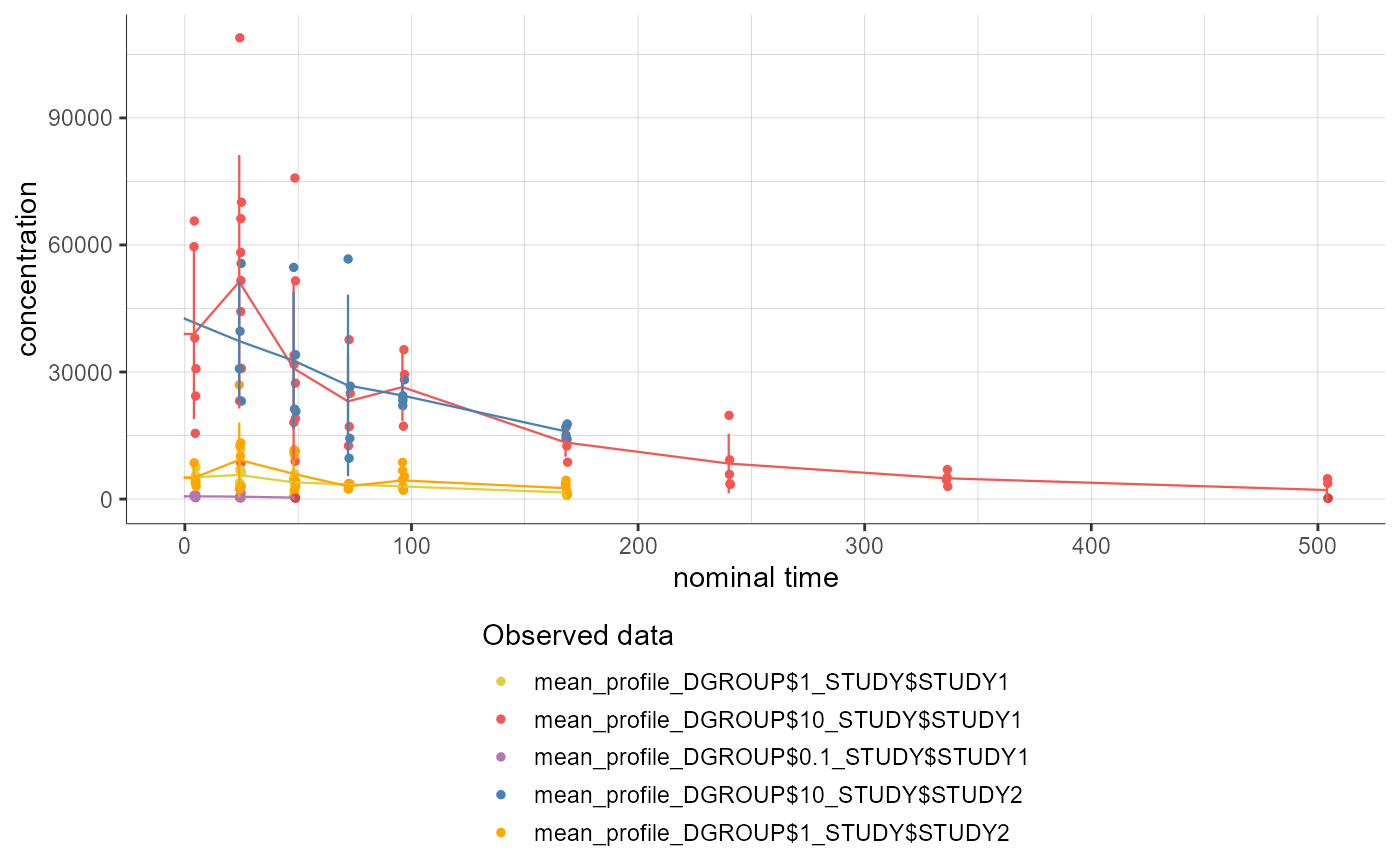
# displaying only the averaged profiles, without individual dots and without error bars
plotNCAData(settings=list(dots=F,
error=F))
#> Warning: No shared levels found between `names(values)` of the manual scale and the
#> data's linetype values.