In PKanalix, a dataset is loaded in the Data tab to create a project. After the dataset accepted, it is possible to specify units and filter the dataset, so units and filtering information do not need to be included in the dataset.
The dataset format used in PKanalix is the same across the entire MonolixSuite, so that it is easy to transition between applications. In this format:
-
Each row corresponds to one individual and one time point.
-
Each row can include a single measurement (also called observation), or a dose amount, or both a measurement and a dose amount.
-
Dosing information should be indicated for each individual in a specific column, even if it is the same for all individuals.
-
There are no restrictions on headers(column names), but there can be only one header row.
-
Different types of information (dose, observation, covariate, etc.) are recorded in different columns, which must be tagged with a column type (see below).
-
Multiple types of observations are all specified in the same column and identified by a separate observation id or occasion column (see below) .
If your dataset is not already in this format, in most cases it is possible to format it in a few steps with the data formatting module, to incorporate missing information (e.g., dose amounts or covariates) or combine columns (e.g., observations in separate columns).
Overview of common column-types
A dataset typically contains at least the following columns: ID (mandatory), TIME (mandatory), OBSERVATION (mandatory), AMOUNT (optional). The main rules for interpreting the dataset are:
-
Missing values should be represented by a period (dot) (e.g., the dose AMOUNT column in a row for a measurement).
-
There are no restrictions on headers(column names), but there can be only one header row. Each column must be assigned to one of the available column-types (columns can be ignored).
The full list of column-types is available at the end of this page and a detailed description is given on the data set documentation website.
The most common column types in addition to ID, TIME, OBSERVATION and AMOUNT are:
-
For IV infusion data, either INFUSION RATE or INFUSION DURATION is required.
-
For steady-state data, either an INTERDOSE INTERVAL column is present, or the interdose interval tau must be specified in the NCA settings.
-
If a dose and a measurement occur at the same time, they can be specified on the same row or on two separate rows.
-
Sort and carry variables can be defined using the OCCASION, CATEGORICAL COVARIATE, and CONTINUOUS COVARIATE column-types.
-
BLQ data are defined using the CENSORING and LIMIT column-types.
In addition to the typical cases presented above, a few additional column-types may also be useful:
-
ignoring a row: This is possible with the IGNORED OBSERVATION (ignores the measurement only) or IGNORED LINE (ignores the entire row, including regressor and dose information) column-types. However, it is more convenient to filter rows of your dataset without modifying it by using filters (available once your dataset is accepted).
-
working with several types of observations: If several observation types are present in the dataset (for example parent and metabolite), all measurements should still appear in the same OBSERVATION column, and another column should be used to distinguish the observation types. If this is not the case in your dataset, data formatting can merge several observation columns. To run NCA for several observations at the same time for the same id, tag the column with the observation types as OCCASION. To run CA for several observations with a model including several outputs (for example PK and PD), tag the column listing the observation types as OBSERVATION ID. In this case, only one observation type will be available for NCA. It can be selected in the “NCA” section to perform the calculations.
-
working with several types of administrations: a single data set can contain different types of administrations, for instance, IV bolus and extravascular, distinguished using the ADMINISTRATION ID column-type. The setting “administration type” in “Tasks>Run” can be chosen separately for each administration id, and the appropriate parameter calculations will be performed.
Example datasets
Below we show how to specify the dataset for several common situations.
Plasma concentration data
Extravascular
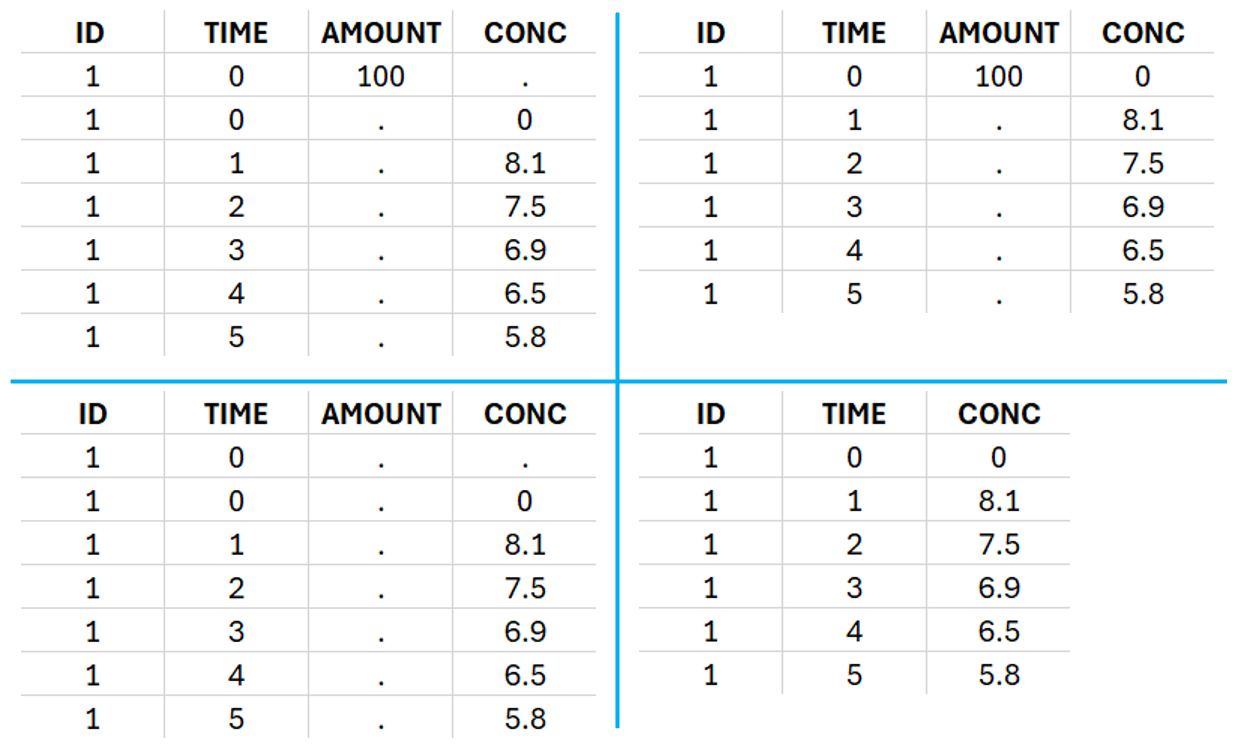
For extravascular administration, the mandatory column-types are ID (individual identifiers as integers or strings), OBSERVATION (measured concentrations), AMOUNT (dose amount administered, mandatory for 2023 and previous versions) and TIME (time of dose or measurement). Starting with version 2024, datasets that do not contain an AMOUNT column or no information in the AMOUNT column are accepted for single dose or multiple dose administrations (bottom row in figure below). If there are concentration measurements and doses at the same time, they can be specified on the same row or separately (top row in figure below).
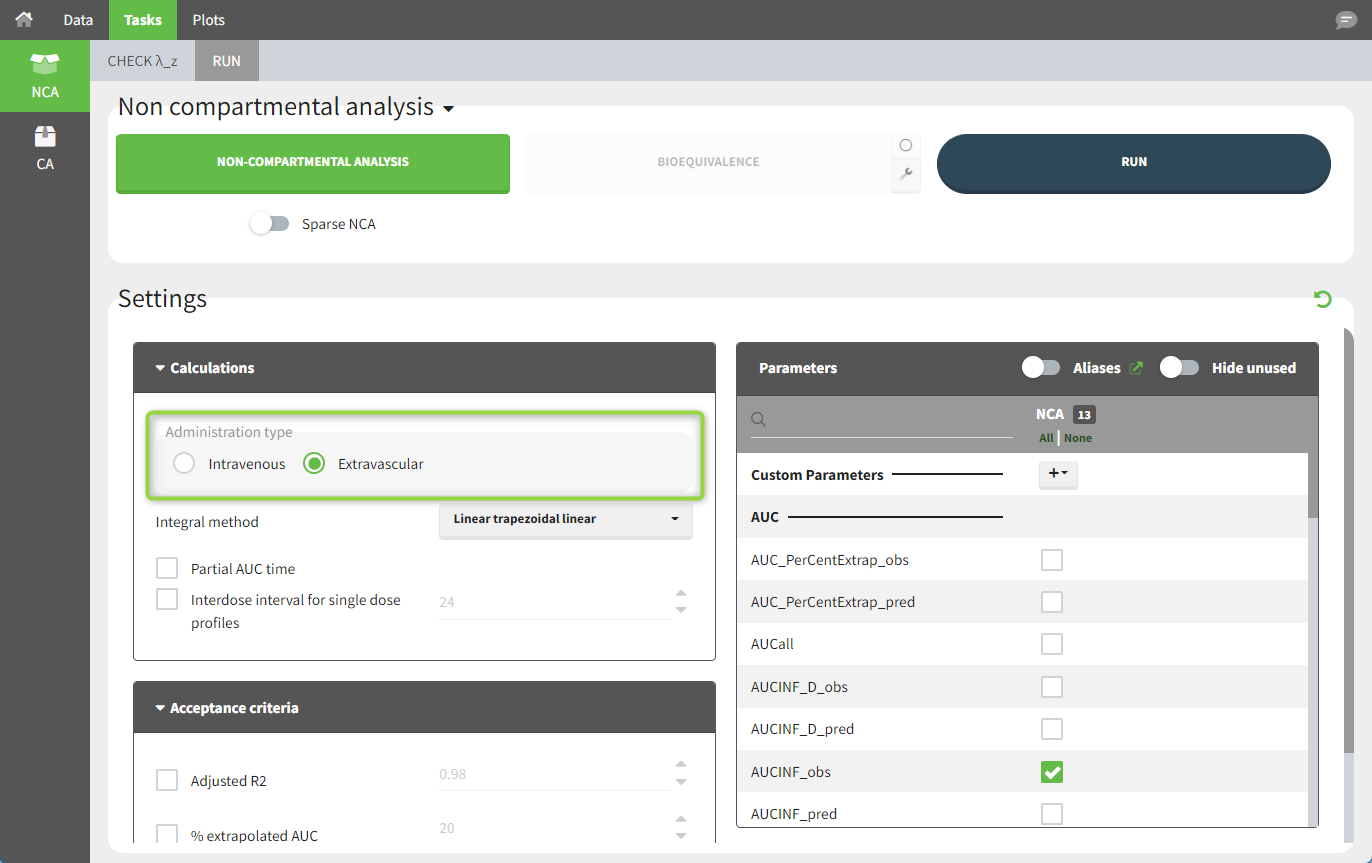
To distinguish an extravascular administration from an IV bolus administration, in “Tasks>Run” the administration type must be set to “extravascular.”
If no measurement is recorded at the time of the dose, a concentration of zero is added for single dose data, the minimum concentration observed during the dose interval for steady-state data.
Example:
-
demo project_extravascular.pkx
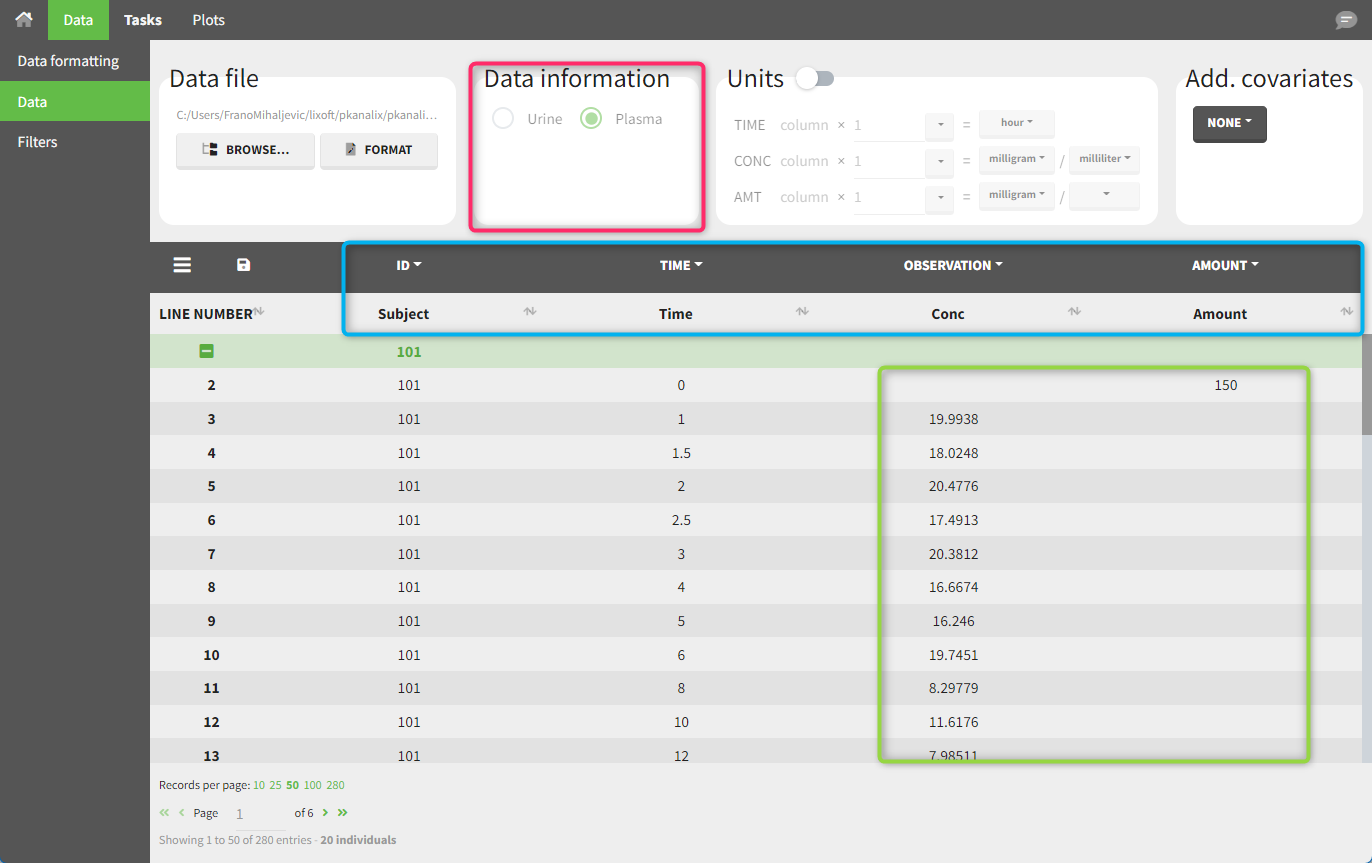
This data set records the drug concentration measured after single oral administration of 150 mg of the drug in 20 patients. For each individual, the first row records the dose (in the “Amount” column tagged as AMOUNT column-type) while the following rows record the measured concentrations (in the “Conc” column tagged as OBSERVATION). Cells in the “Amount” column for measurement rows contain a period, and similarly for the concentration column. The column containing the times of measurements or doses is tagged with the TIME column-type and the subject identifiers, which we will use as a sort variable, are tagged as ID. Use the OCCASION column-type if more sort variables are needed. After accepting the dataset, the data is automatically assigned as “Plasma.”
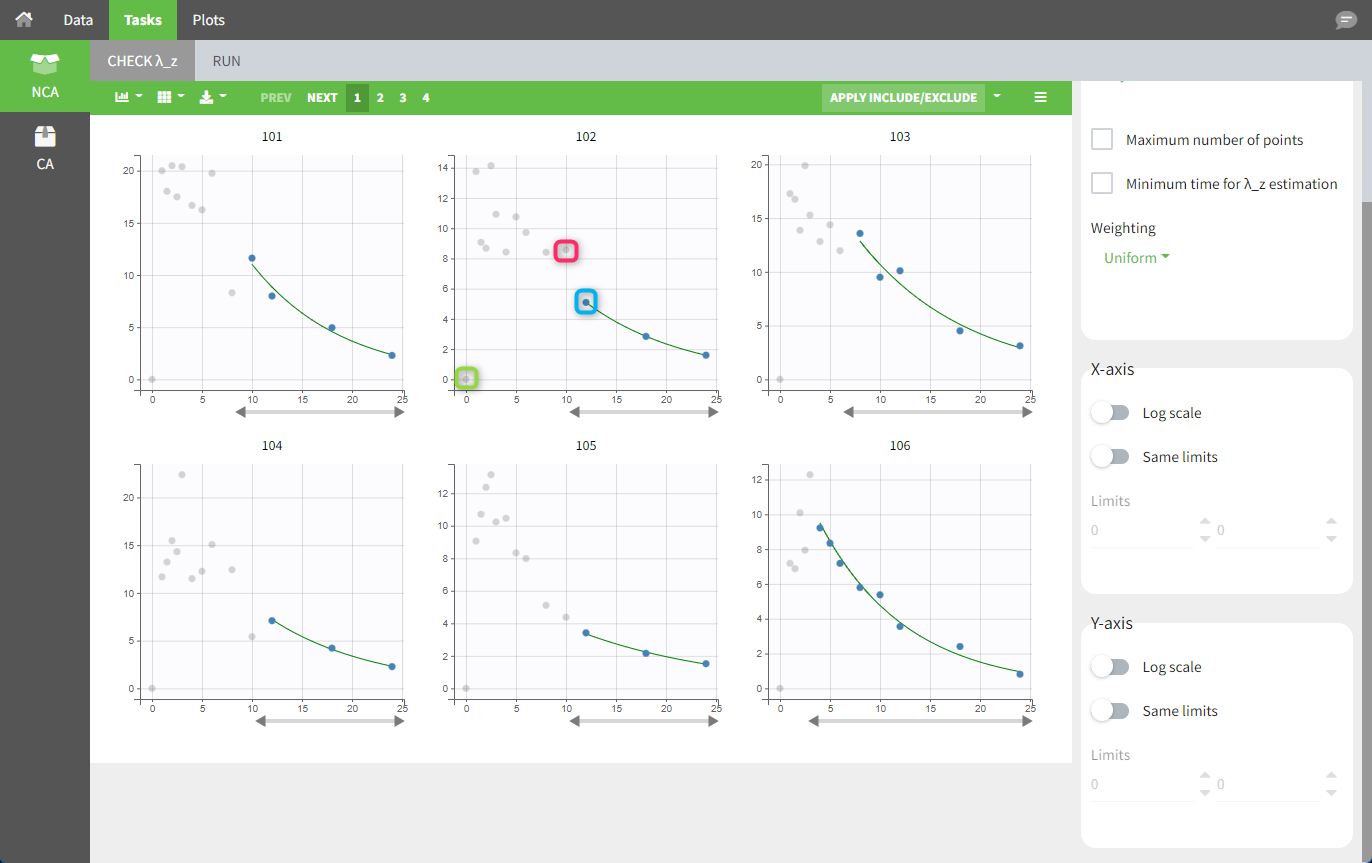
In the “Tasks/Run” tab, the user must indicate that this is extravascular data. In the “Check lambda_z” tab, data points as well as added points, such as a zero concentration at the dose time (only displayed with a linear scale for the y-axis), are shown. Points included in the λz calculation are highlighted in blue.
After running the NCA analysis, PK parameters relevant to extravascular administration are displayed in the “Results” tab.
IV infusion
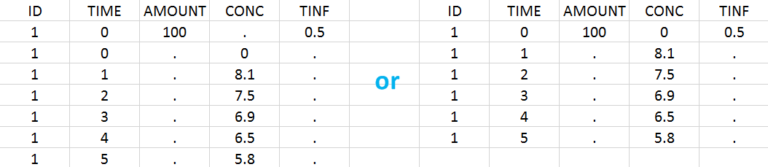
Intravenous infusions are indicated in the data set via the presence of an INFUSION RATE or INFUSION DURATION column-type, in addition to the ID (individual identifiers as integers or strings), OBSERVATION (measured concentrations), AMOUNT (dose amount administered, mandatory for 2023 and previous versions) and TIME (time of dose or measurement). The infusion duration (or rate) can be identical or different among individuals. Starting with version 2024, datasets that do not contain an AMOUNT column or no information in the AMOUT column are accepted for single dose or multiple dose administrations. If there are concentration measurements and doses at the same time, they can be specified on the same row or separately (see figure below).
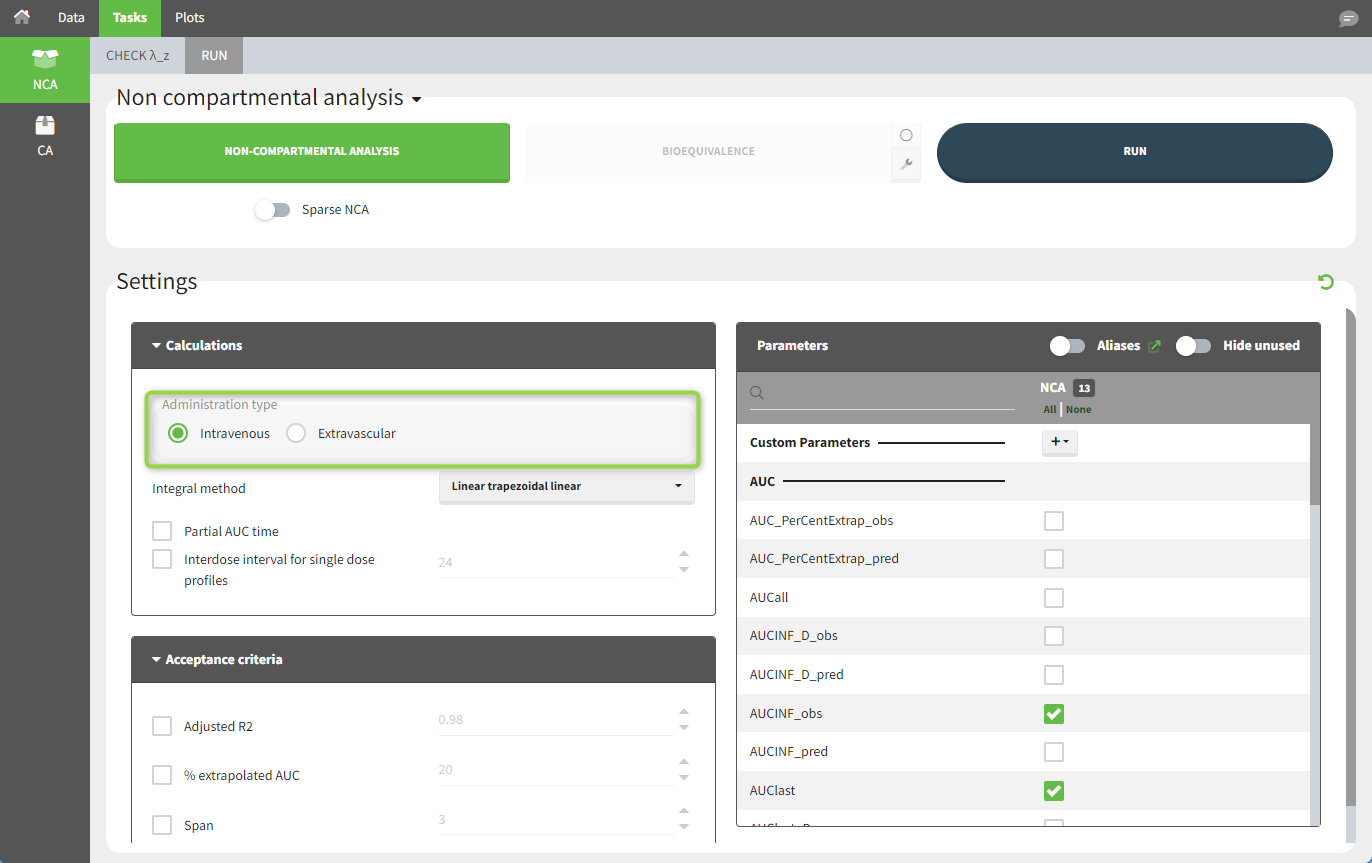
In “Tasks>Run” the administration type must be set to “intravenous”.
If no measurement is recorded at the time of the dose, a concentration of zero is added for single dose data and the minimum concentration observed during the dose interval is added for steady-state data.
Example:
-
demo project_ivinfusion.pkx:
In this example, the patients receive an IV infusion taking 3 hours. The infusion duration is recorded in the column named “TINF” in this example, and tagged as INFUSION DURATION.
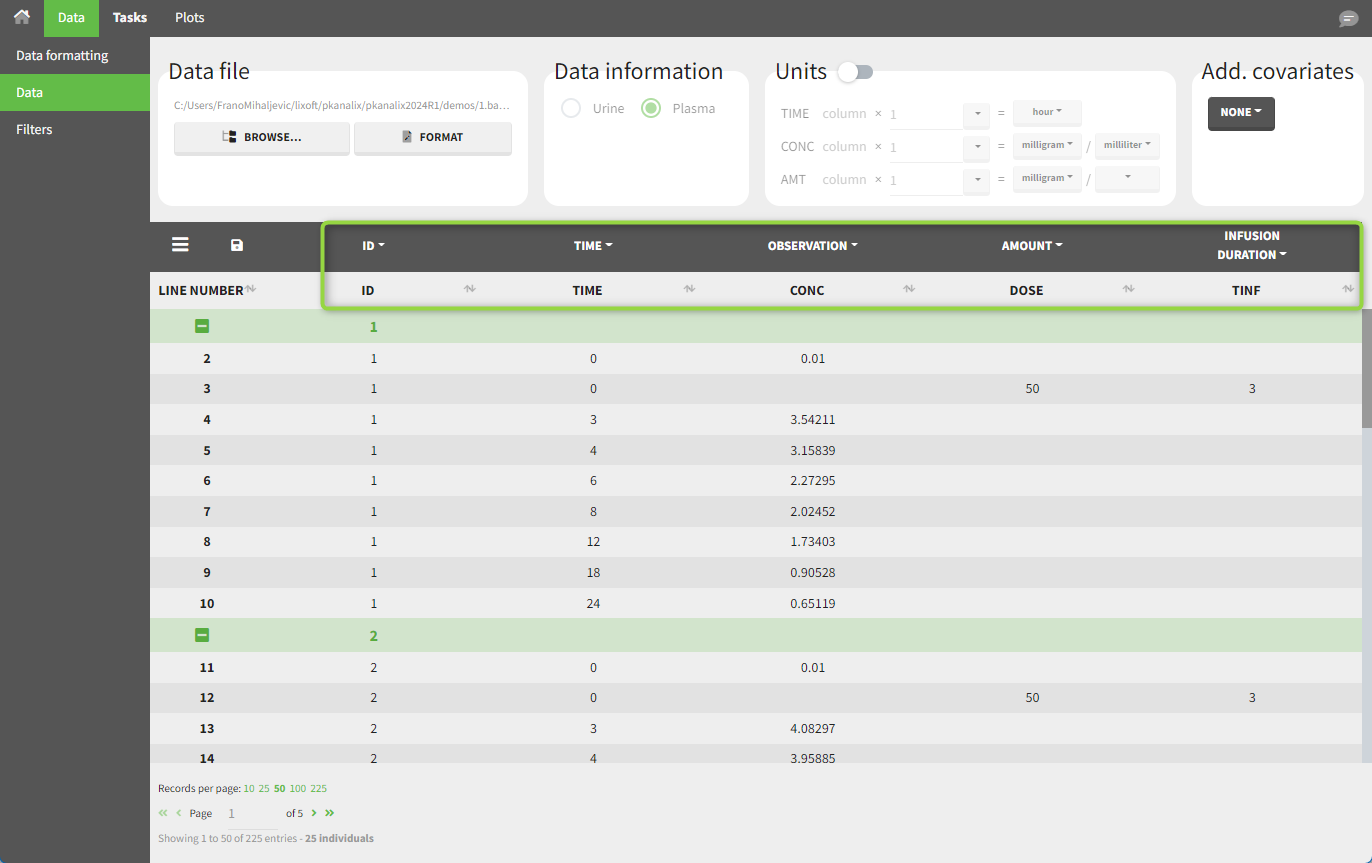
In the “Tasks/Run” tab, the user must indicate that this is intravenous data.
IV bolus
For IV bolus administration, the mandatory column-types are ID (individual identifiers as integers or strings), OBSERVATION (measured concentrations), AMOUNT (dose amount administered, mandatory for 2023 and previous versions) and TIME (time of dose or measurement). Starting with version 2024, datasets that do not contain an AMOUNT column or no information in the AMOUNT column are accepted for single dose or multiple dose administrations (bottom row in figure below). If there are concentration measurements and doses at the same time, they can be specified on the same row or separately (top row in figure below).
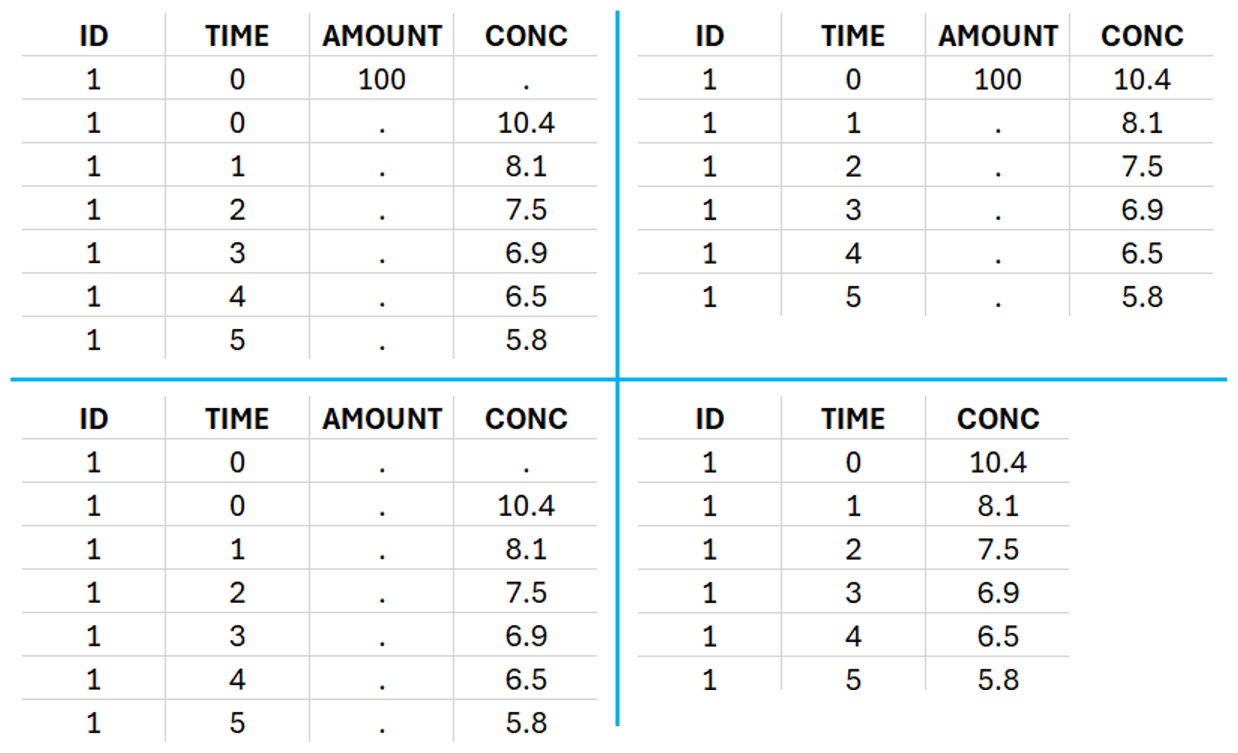
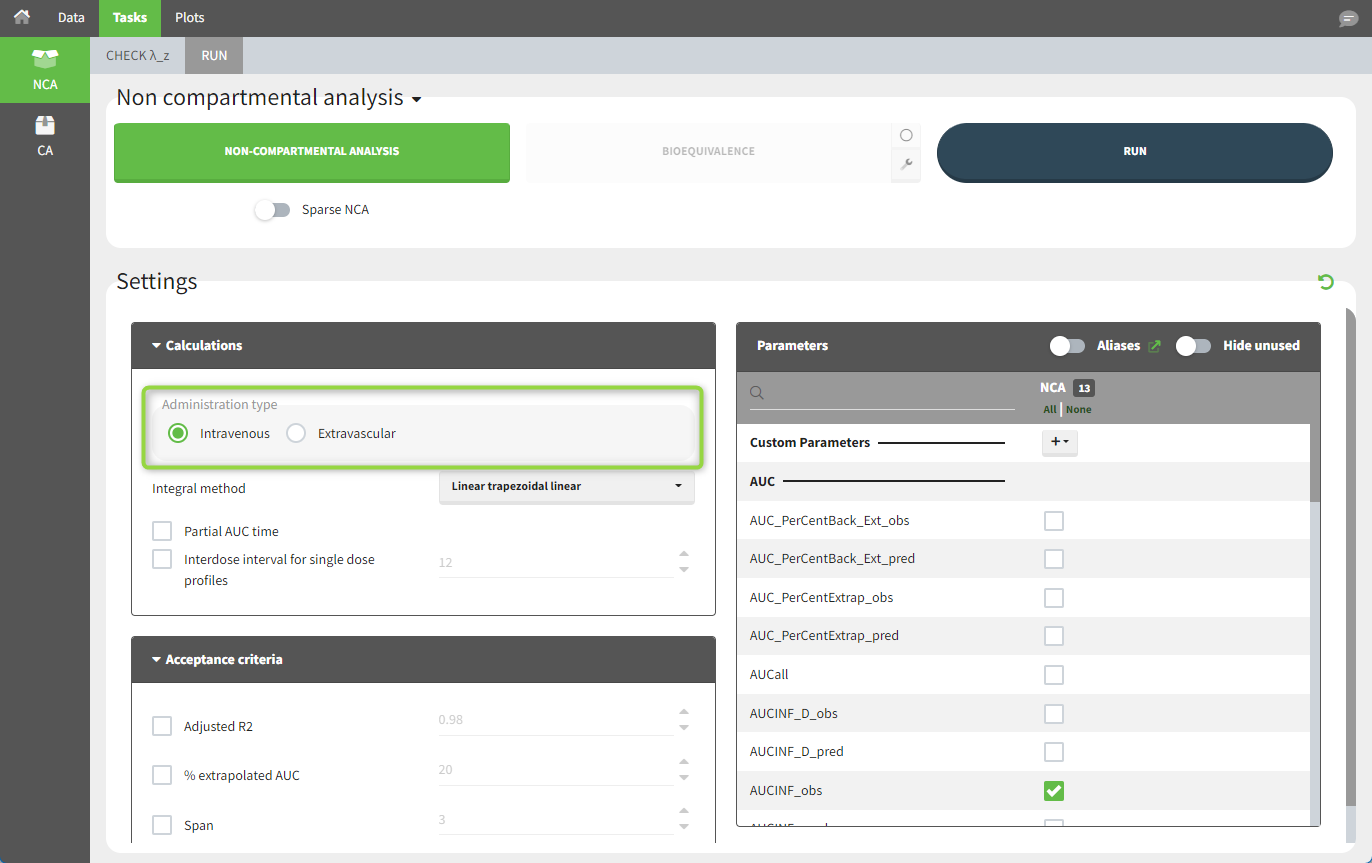
To distinguish the IV bolus from the extravascular case, in “Tasks>Run” the administration type must be set to “intravenous”.
If no measurement is recorded at the time of the dose, the concentration of at time zero is extrapolated using a log-linear regression of the first two data points, or is taken to be the first observed measurement if the regression yields a slope >= 0. See the calculation details for more information.
Example:
-
demo project_ivbolus.pkx:
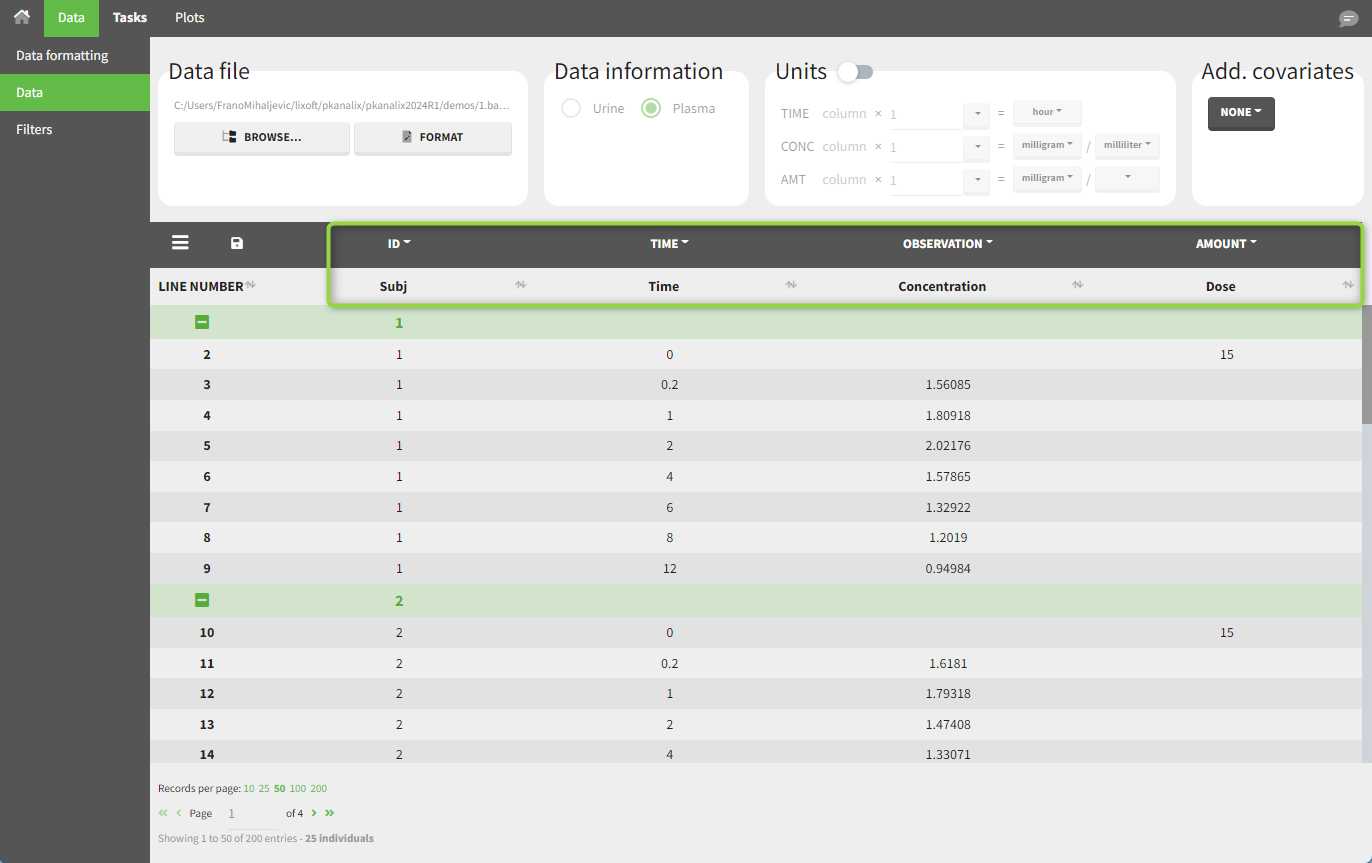
In this data set, 25 individuals have received an IV bolus and their plasma concentration have been recorded over 12 hours. For each individual (indicated in the column “Subj” tagged as ID column-type), we specify the dose amount in a column “Dose,” tagged as the AMOUNT column-type. The measured concentrations are tagged as OBSERVATION and the times as TIME. Use the OCCASION column-type if more sort variables are needed in addition to ID. After accepting the dataset, the data is automatically assigned as “plasma.”
In the “Tasks/Run” tab, the user must indicate that this is intravenous data. In the “Check lambda_z” tab, both measurements originally present in the data and added data points, such as the C0 at the dose time, are shown. Data points included in the λz calculation are highlighted in blue.
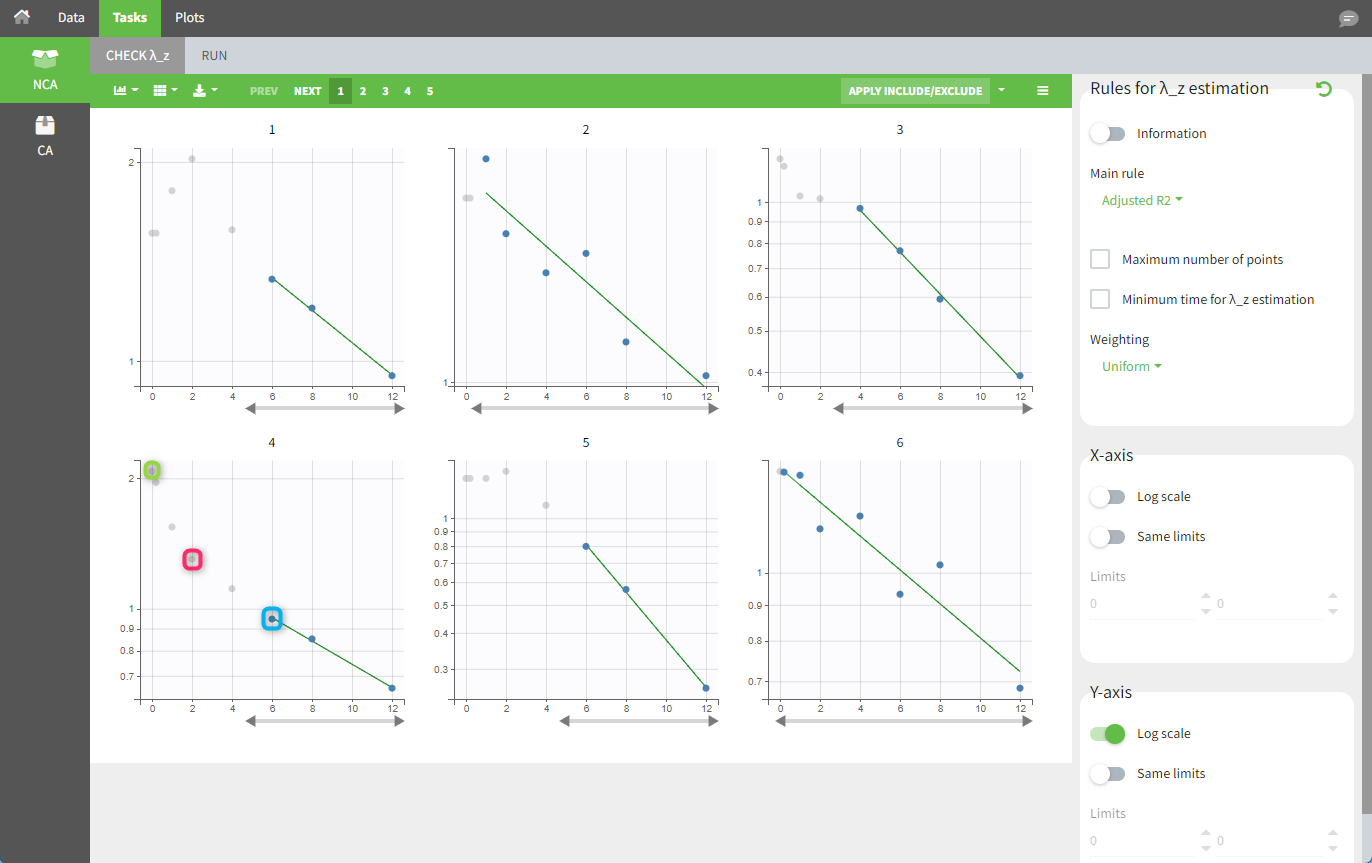
After running the NCA analysis, PK parameters relevant to IV bolus administration are displayed in the “Results” tab.
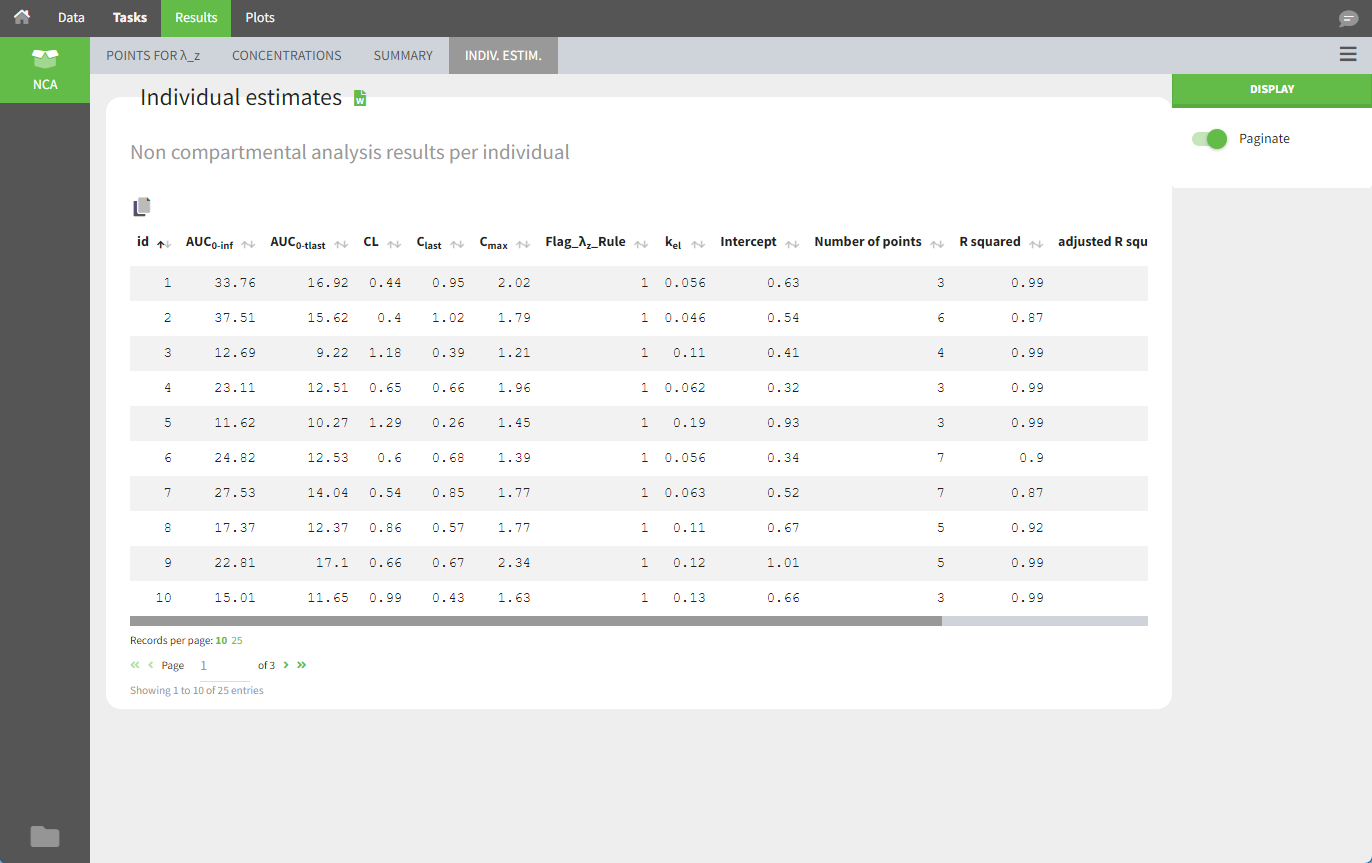
Steady-state
Starting with version 2024, the interdose interval tau used to calculate NCA parameters specific to steady-state can be specified either in the dataset column tagged as INTERDOSE INTERVAL, or directly as part of the NCA settings.
In version 2023 and previous, it was necessary to specify steady-state using the STEADY STATE column-type: SS=1 indicates that the individual is already at steady-state when receiving the dose. This implicitly assumes that the individual has received many doses before this one. SS=0 or ‘.’ indicates a single dose. Starting with version 2024, the SS column is still accepted but no longer mandatory.
The dosing interval (also called tau) is specified in the INTERDOSE INTERVAL column on the rows defining the doses, or as part of the NCA settings.
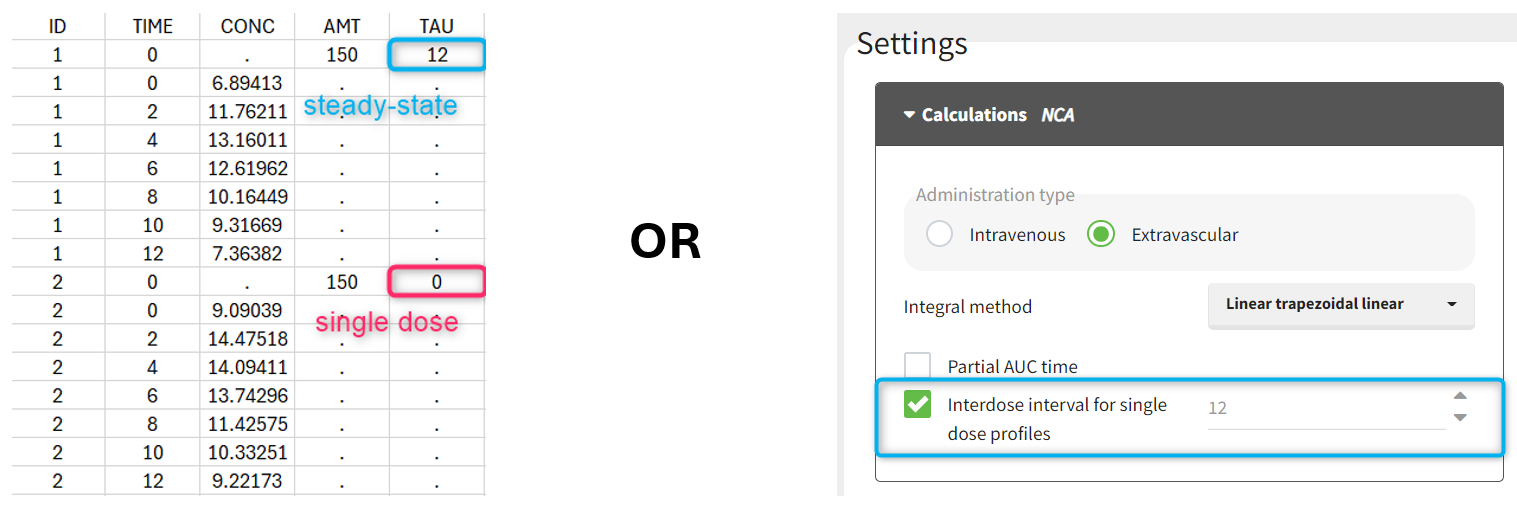
Steady state calculation formulas are applied for individuals with a dose with INTERDOSE INTERVAL = <double>. A data set can contain individuals which are at steady-state and some which are not. If the NCA setting “Interdose interval for single dose profiles” is selected, steady-state parameters are calculated for all individuals.
If no measurement is recorded at the time of the dose, the minimum concentration observed during the dose interval is added at the time of the dose for extravascular and infusion data. For IV bolus data, a regression using the two first data points is performed. Only measurements between the dose time and dose time + interdose interval will be used.
Examples:
-
demo project_steadystate.pkx:
In this example, the individuals are already at steady-state when they receive the dose. This is indicated in the dataset via the column “SteadyState” tagged as the STEADY STATE column-type, which contains a “1” on rows recording doses. The interdose interval is specified on those same rows in the column “tau” tagged as INTERDOSE INTERVAL. When accepting the dataset, a “Settings” section appears, where one can define the number of steady-state doses. This information is relevant when exporting to Monolix, but not directly used in PKanalix.
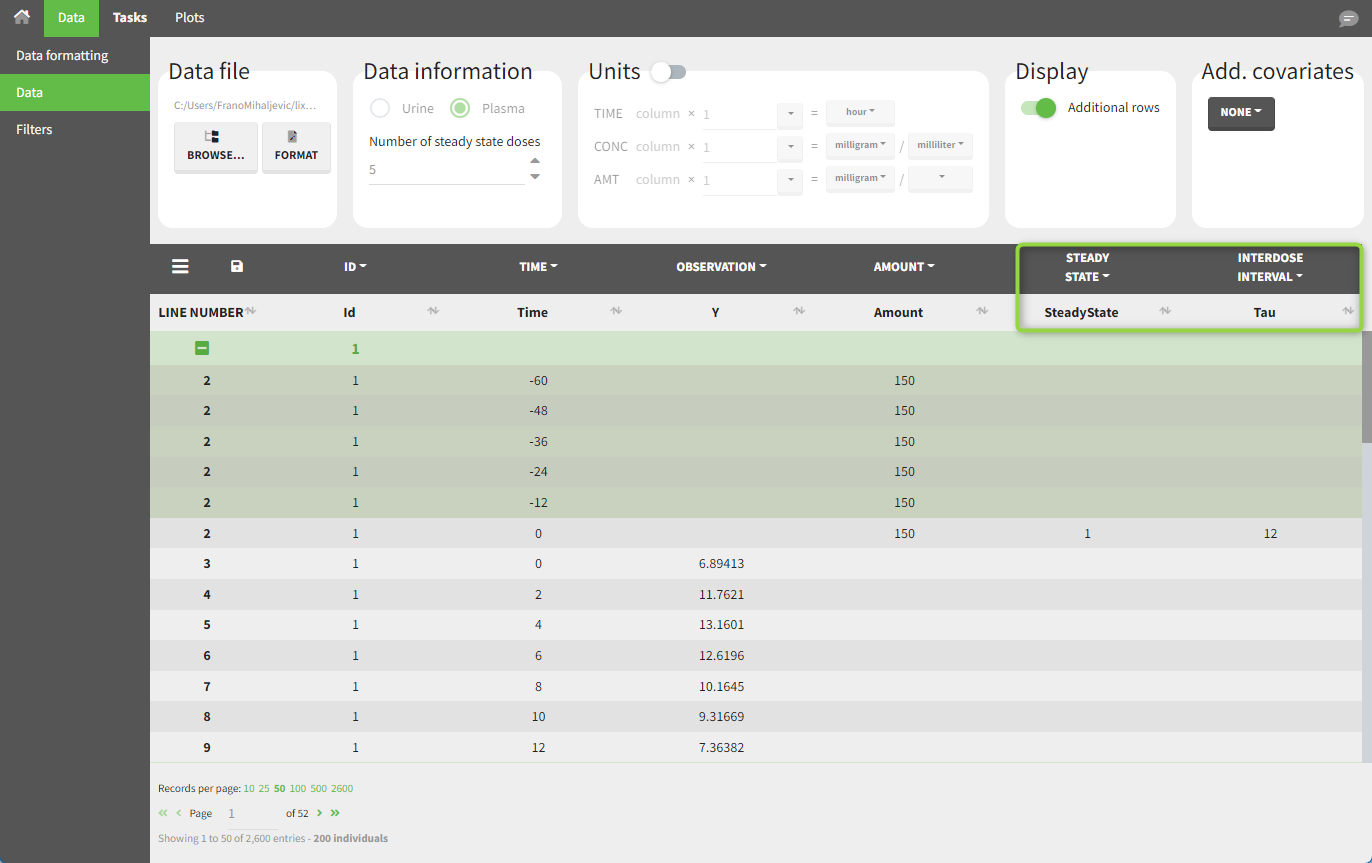
After running the NCA estimation task, steady-state specific parameters are displayed in the “Results” tab.
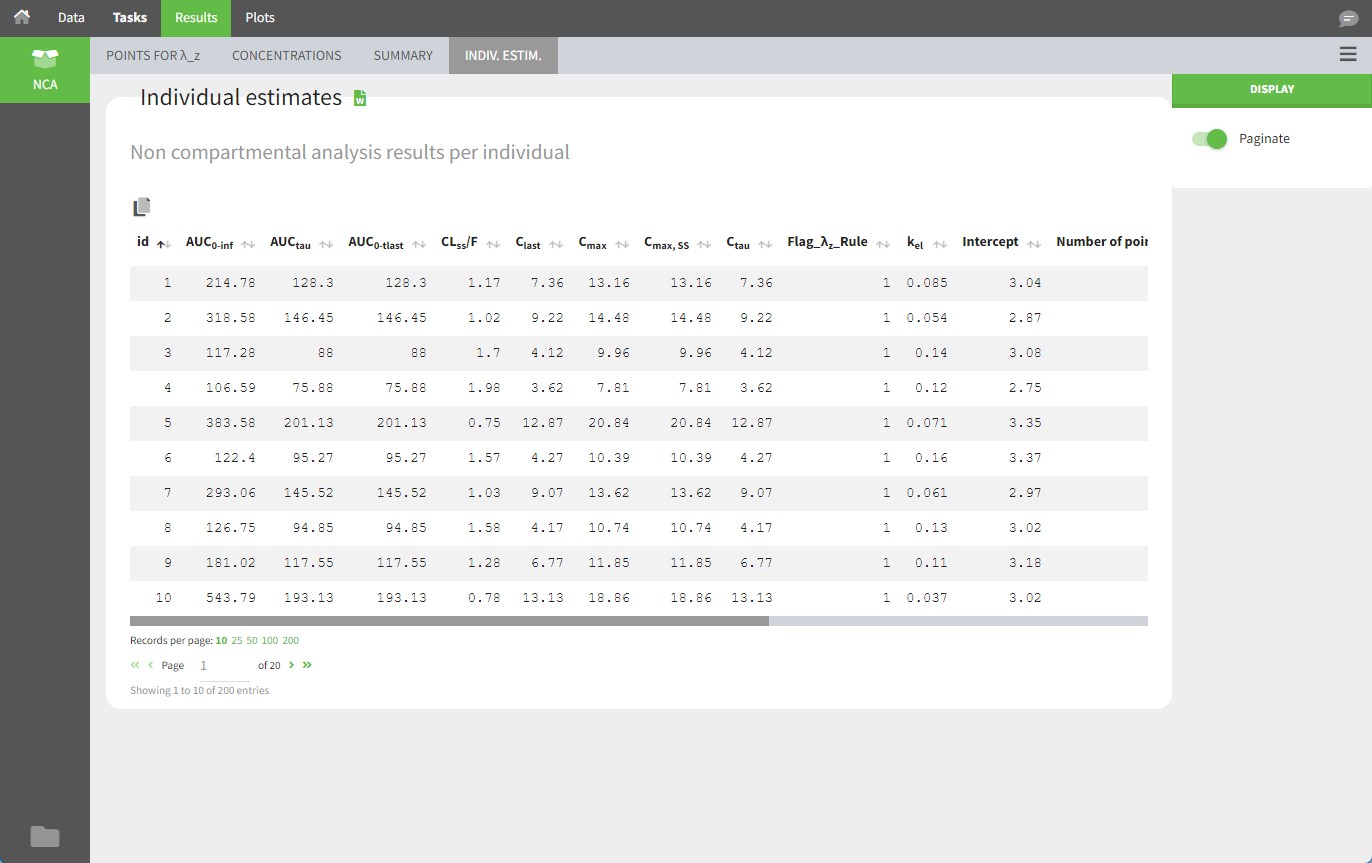
BLQ data
Below the limit of quantification (BLQ) data can be recorded in the data set using the CENSORING column:
-
“0” indicates that the value in the OBSERVATION column is the measurement.
-
“1” indicates that the observation is BLQ.
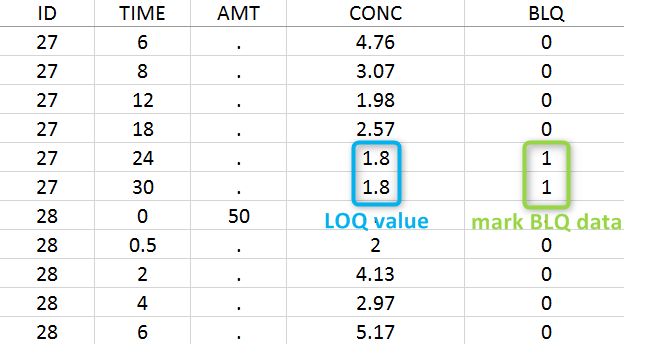
The lower limit of quantification (LOQ) must be indicated in the OBSERVATION column when CENSORING = “1”. Note that strings are not allowed in the OBSERVATION column (except periods). A different LOQ value can be used for each BLQ measurement.
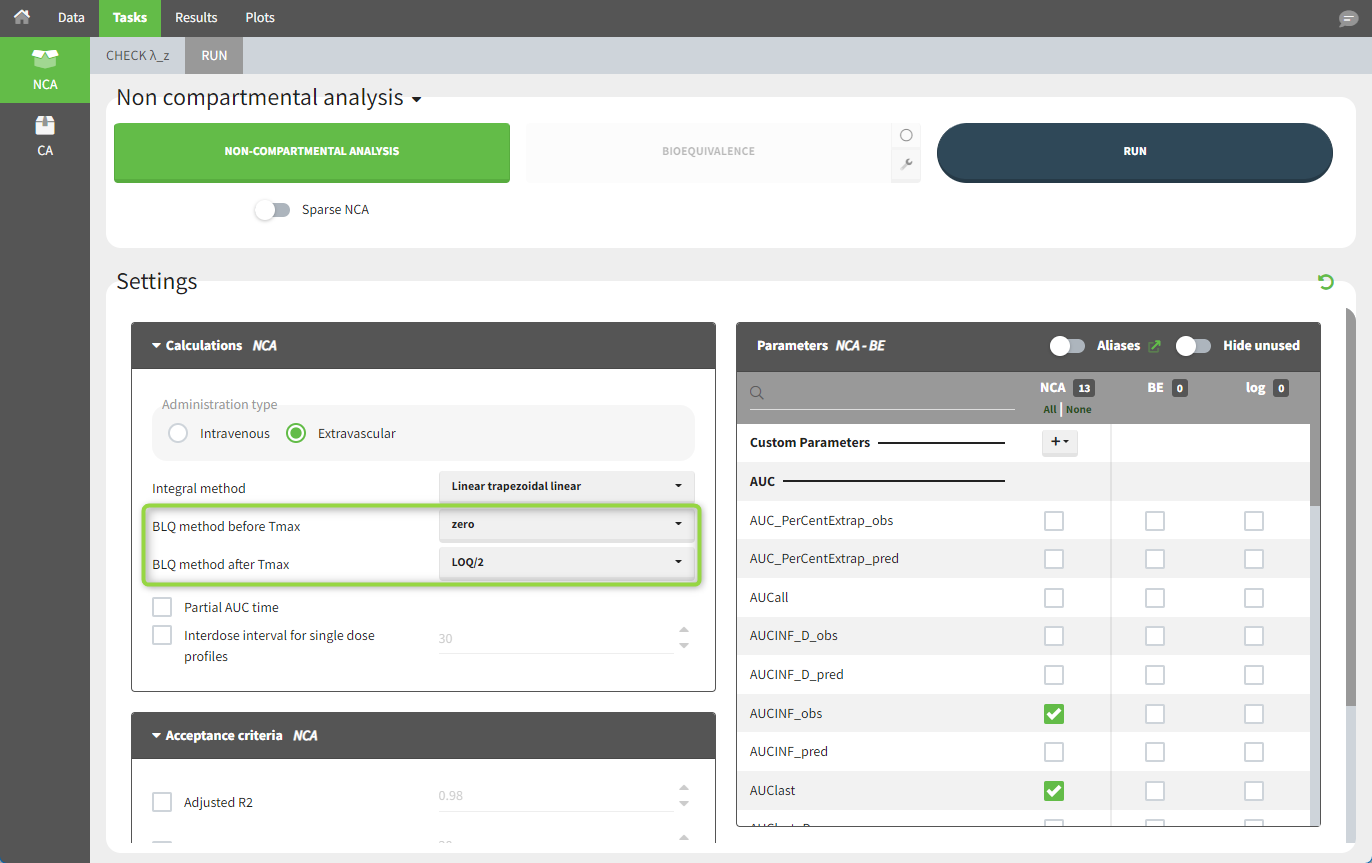
When performing an NCA analysis, the BLQ data before and after the Tmax are can be handled differently. They can be replaced by:
-
zero
-
the LOQ value
-
the LOQ value divided by 2
-
or considered as missing
For a CA analysis, the same options are available, but no distinction is made before and after Tmax. Once replaced, the BLQ data are treated as any other observation.
A LIMIT column can be added to record the other limit of the interval (generally zero). This value will not be used by PKanalix but can facilitate the transition from an NCA/CA analysis in PKanalix to a population model in Monolix.
To easily transform BLQ data in a dataset that has BLQ tags in the observation column, you can use Data formatting.
Examples:
-
demo project_censoring.pkx: two studies with BLQ data with two different LOQ values
In this dataset, the measurements of two different studies (indicated in the STUDY column, tagged as CATEGORICAL COVARIATE in order to be used to stratify results tables and plots) are observed. For the study 101, the LOQ value is 1.8 ug/mL, while it is 1 ug/mL for study 102. The BLQ measurements are marked with a “1” in the BLQ column, which is tagged as CENSORING. The LOQ values are specified for each BLQ measurement in the CONC column of measurements, tagged as OBSERVATION.
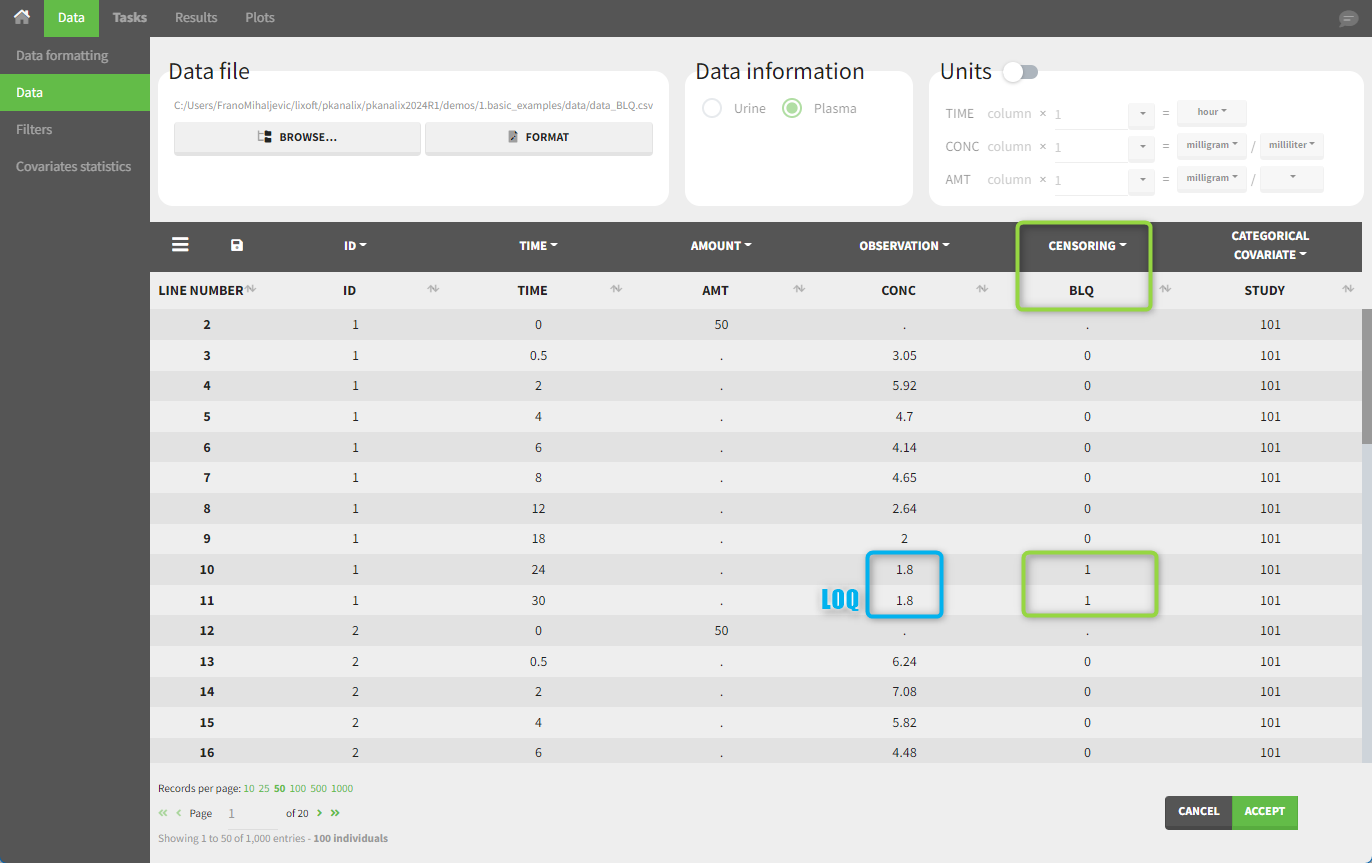
For the NCA analysis, in the “Task>NCA>Run” tab, the user can choose how to handle the BLQ measurements. For the BLQ measurements before and after the Tmax, the BLQ measurements can be considered as missing (as if this row in the dataset did not exist), or replaced by zero (the default before Tmax), the LOQ value, or half the LOQ value (the default after Tmax). In the “Check lambda_z” tab, the BLQ measurements are shown in red and displayed according to their replacement value.
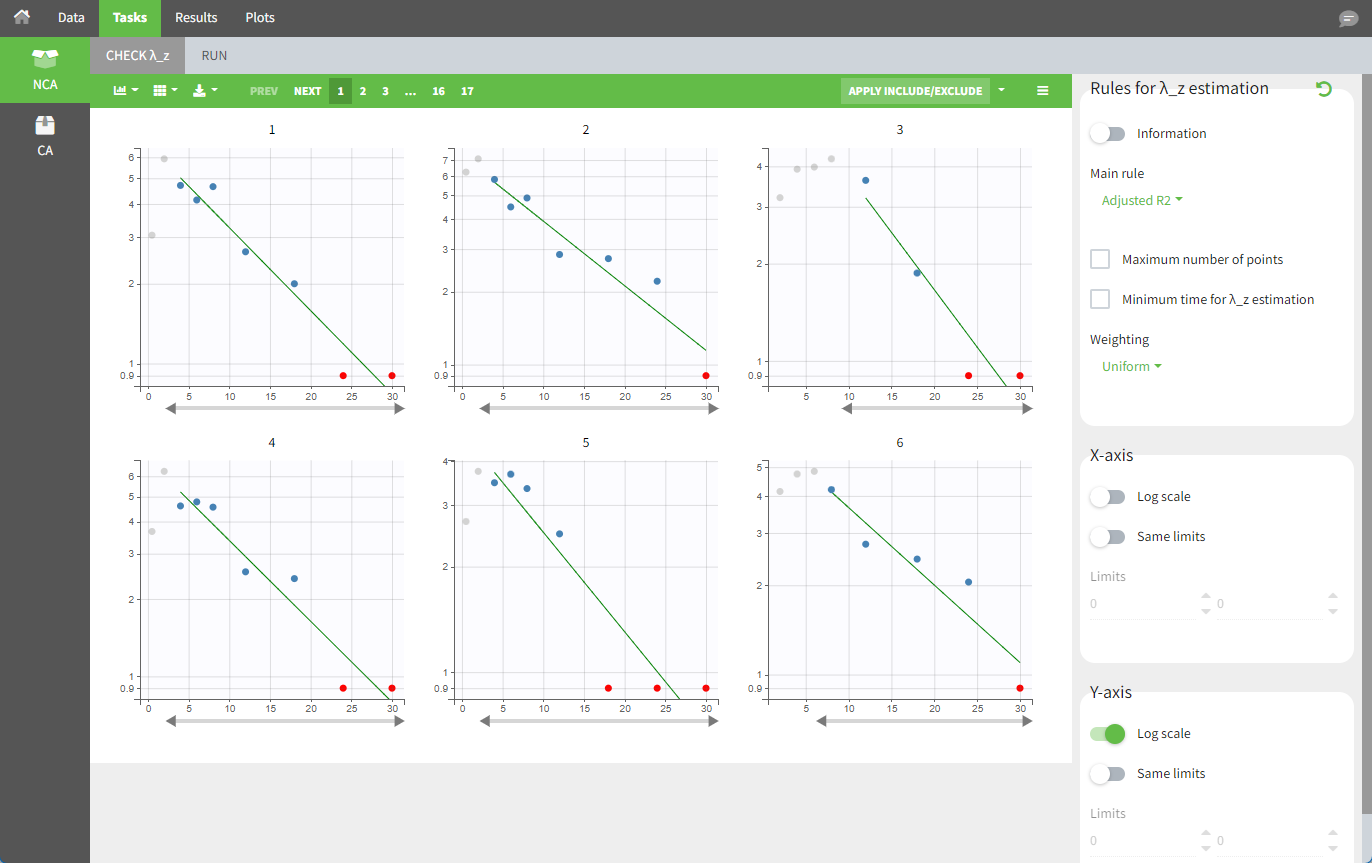
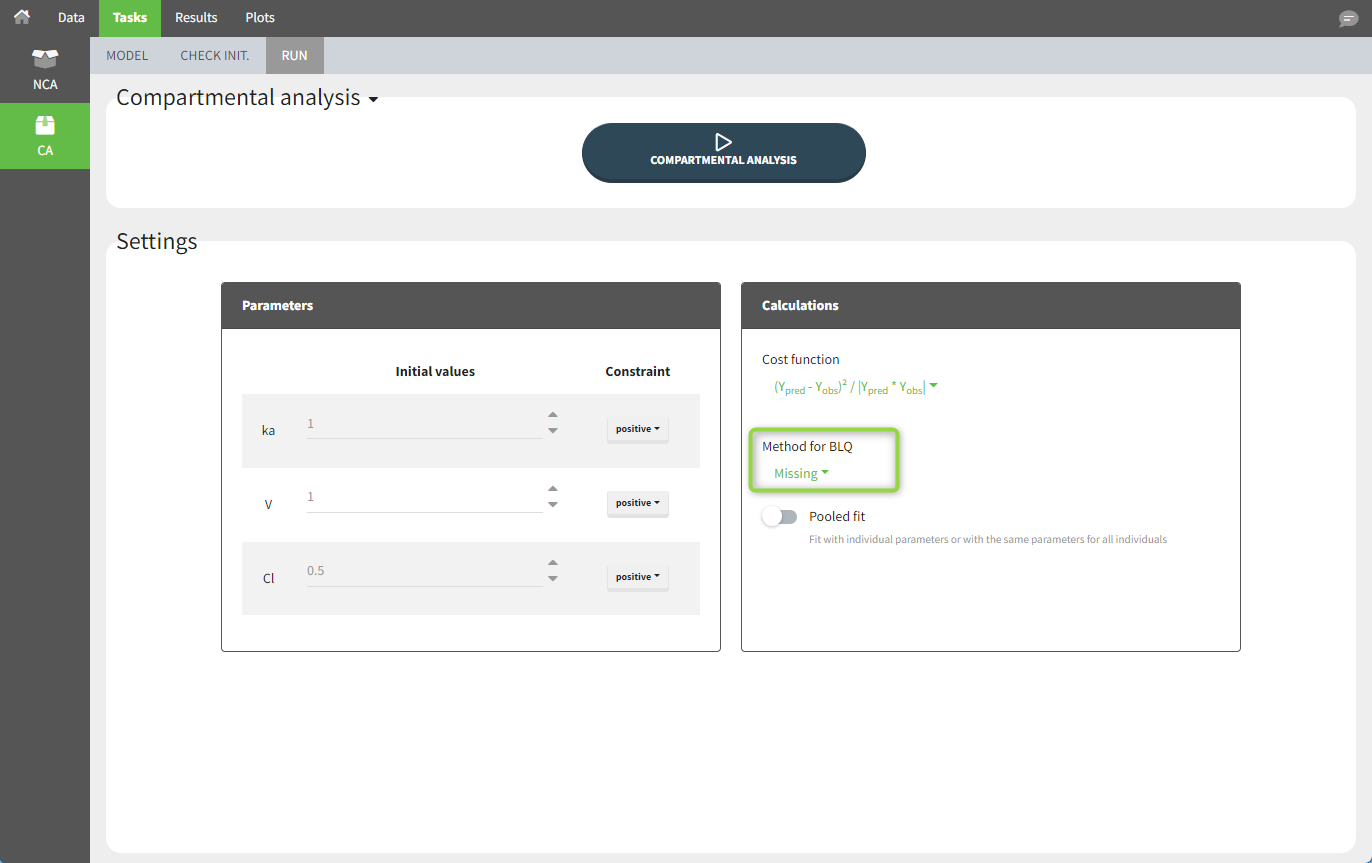
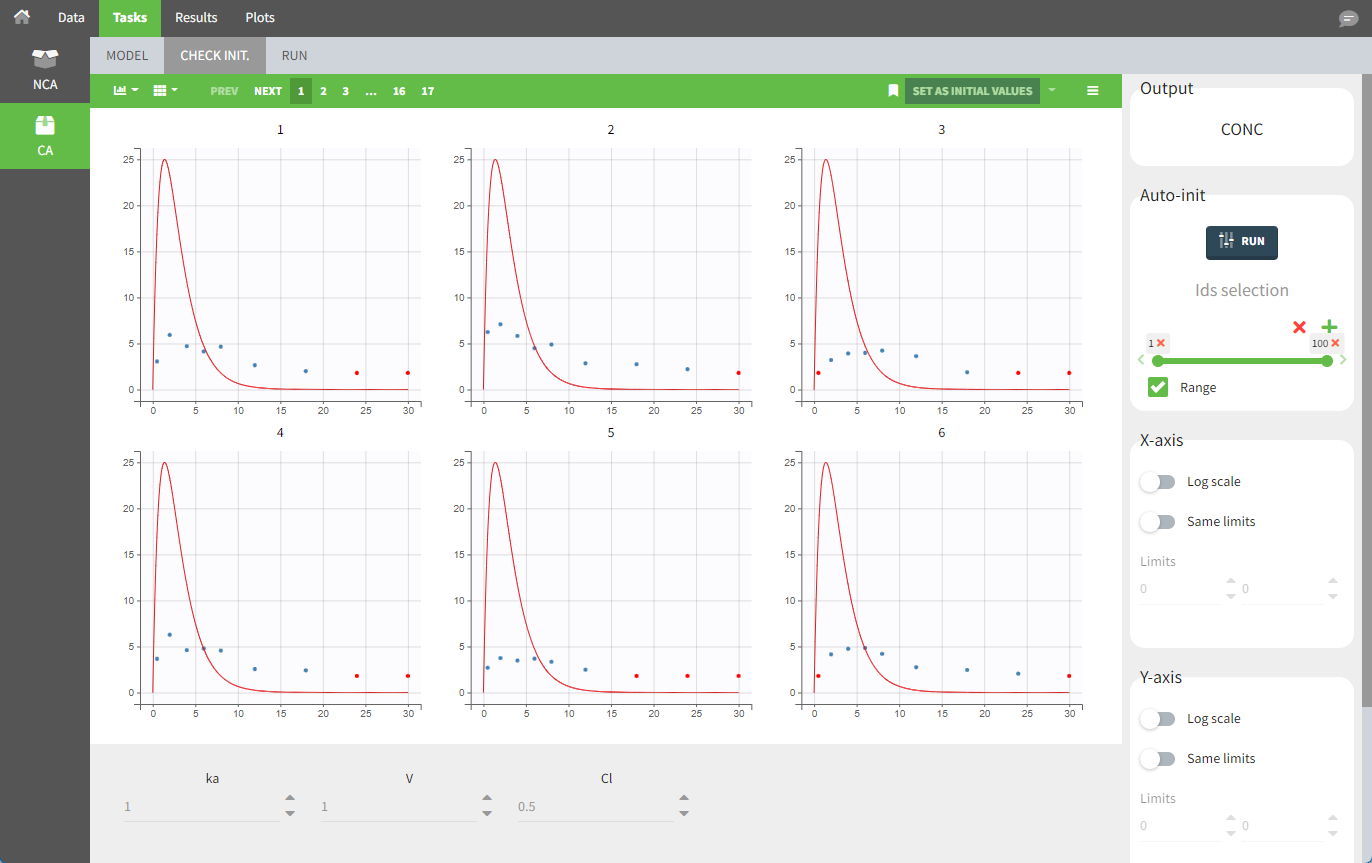
For the CA analysis, the replacement value for all BLQ measurements can be chosen in the settings of the “Run” tab (the default is missing). In the “Check init.” tab, the BLQ are again displayed in red, at the LOQ value (irrespective of the chosen method for the calculations).
Urine data
To work with urine data, it is necessary to record the time and amount administered, the volume of urine collected for each time interval, the start and end time of the intervals and the drug concentration in a urine sample of each interval. The time intervals must be continuous (no gaps allowed).
In PKanalix, the start and end times of the intervals are recorded in a single column, tagged as TIME column-type. In this way, the end time of an interval automatically acts as start time for the next interval. The concentrations are recorded in the OBSERVATION column. The volume column must be tagged as the REGRESSOR column-type. This general column-type allows easy transitions to the other applications of MonolixSuite. As several REGRESSOR columns are allowed, the user can select which REGRESSOR column should be used as volume in the ‘Data information’ section once urine data is selected. The concentration and volume measured for the interval [t1,t2] are noted on the t2 row. The volume value on the dose row is meaningless, but it cannot be missing ('.'). We thus recommend setting it to zero.
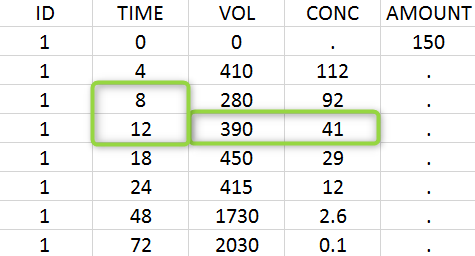
A typical urine dataset has the following structure. A dose of 150 ng has been administered at time 0. The first sampling interval spans from the dose at time 0 to 4h post-dose. During this time, 410 mL of urine have been collected. In this sample, the drug concentration is 112 ng/mL. The second interval spans from 4h to 8h, the collected urine volume is 280 mL and its concentration is 92 ng/mL. The third interval is marked on the figure below: 390 mL of urine have been collected for the interval from 8h to 12h.
The given data is used to calculate the interval midpoints and the excretion rates for each interval. This information is then used to calculate λz and calculate urine-specific parameters. In “Tasks/Check lambda_z” tab, we display the midpoints and excretion rates. However, in the “Plots>Data viewer”, we display the measured concentrations at the end time of the interval.
Example:
-
demo project_urine.pkx: urine PK dataset
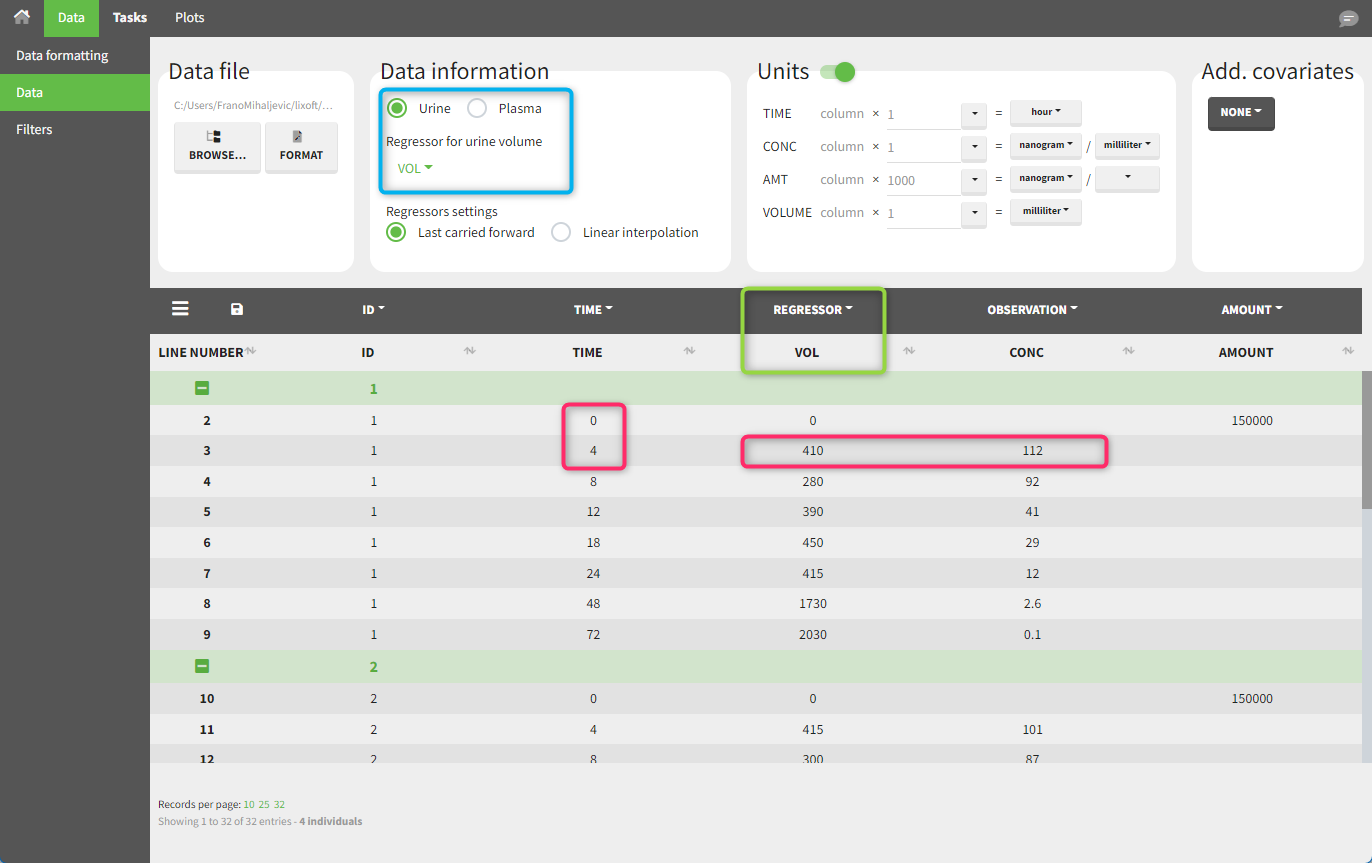
In this urine PK data set, we record the consecutive time intervals in the “TIME” column tagged as TIME. The collected volumes and measured concentration are in the columns “VOL” and “CONC”, respectively tagged as REGRESSOR and OBSERVATION. Note that the volume and concentration are indicated on the row for the interval end time. The volume in the first row (start time of the first interval, as well as dose) is set to zero as it must be a numeric value. This value will not be used in the calculations. Once the dataset is accepted, the observation type must be set to “urine” and the regressor column corresponding to the volume specified.
In the “Tasks>Check lambda_z” tab, the excretion rate are plotted at midpoint times for each individual. The data points selected for the lambda_z calculation works as usual.
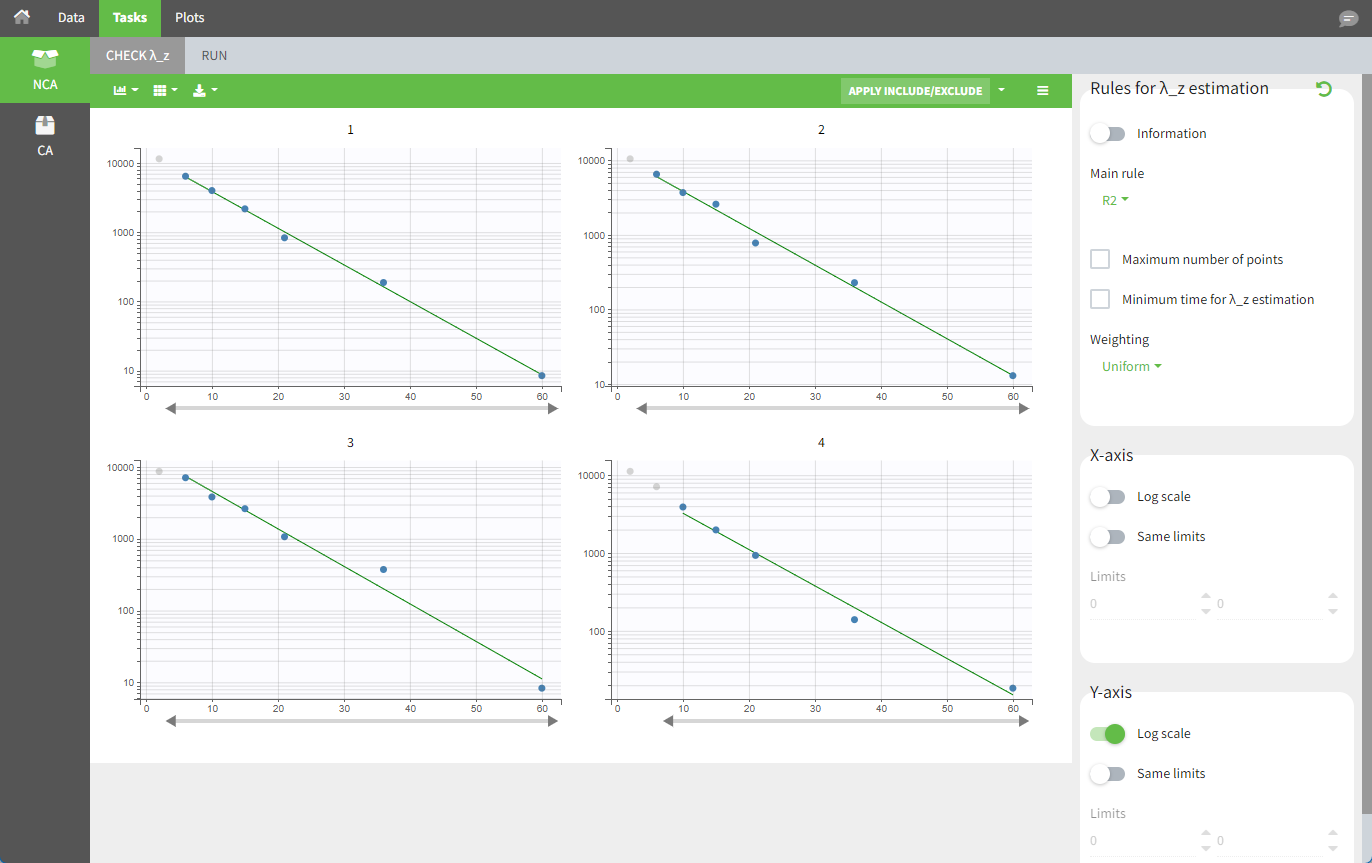
Once the NCA task has run, urine-specific PK parameters are displayed in the “Results” tab.
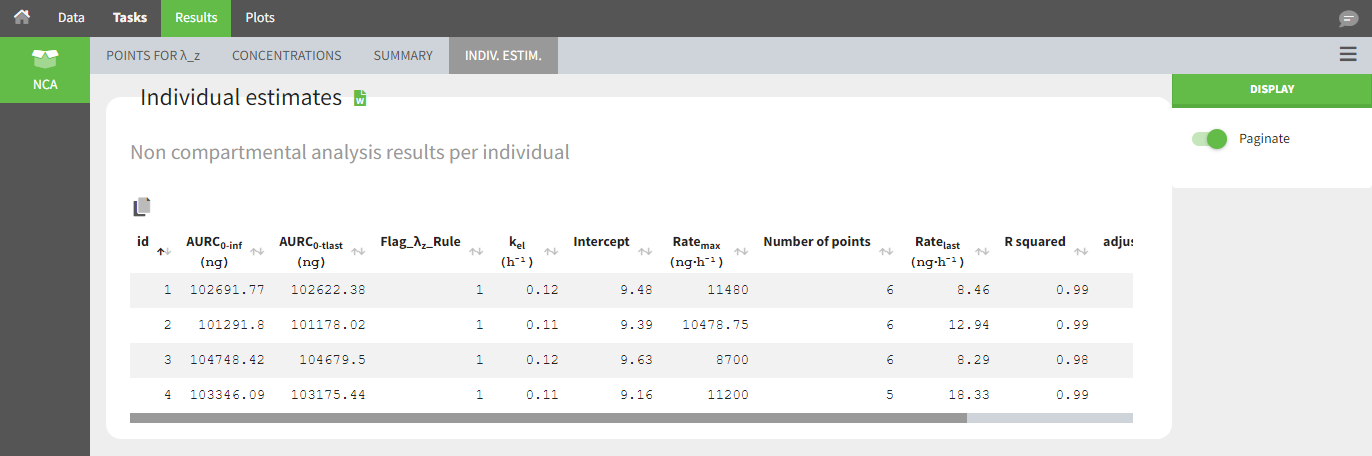
Occasions (“Sort” variables)
The main sort level is by individual indicated by the ID column. Additional sort levels can be specified using one or several OCCASION column(s). OCCASION columns contain integer values that distinguish different time periods for a given individual. The time values can restart at zero or continue when switching from one occasion to the next. The variables that differ among periods, such as the treatment for a crossover study, are tagged as CATEGORICAL or CONTINUOUS COVARIATES (see below). The NCA and CA calculations are performed on each ID-OCCASION combination. Each occasion is considered independent of other occasions (i.e., a washout is applied between each occasion).
Note: occasions columns encoding the sort variables as integers can easily be added to an existing data set using the data formatting module, Excel or R.
With R, the “OCC” column can be added to an existing data frame named “data” with a column “TREAT” with values “ref” and “test” using data$OCC <- ifelse(data$TREAT=="ref", 1, 2).
With Excel, assuming the sort variable is specified in the column E with values “ref” and “test”, use the formula =IF(E2="ref",1,2) to generate the first value of the “OCC” column and then propagate it to the entire column.
Examples:
-
demo project_occasions1.pkx: a crossover study with two treatments
The subject column is tagged as ID, the treatment column as a CATEGORICAL COVARIATE and an additional column encoding the two periods with integers “1” and “2” as an OCCASION column.
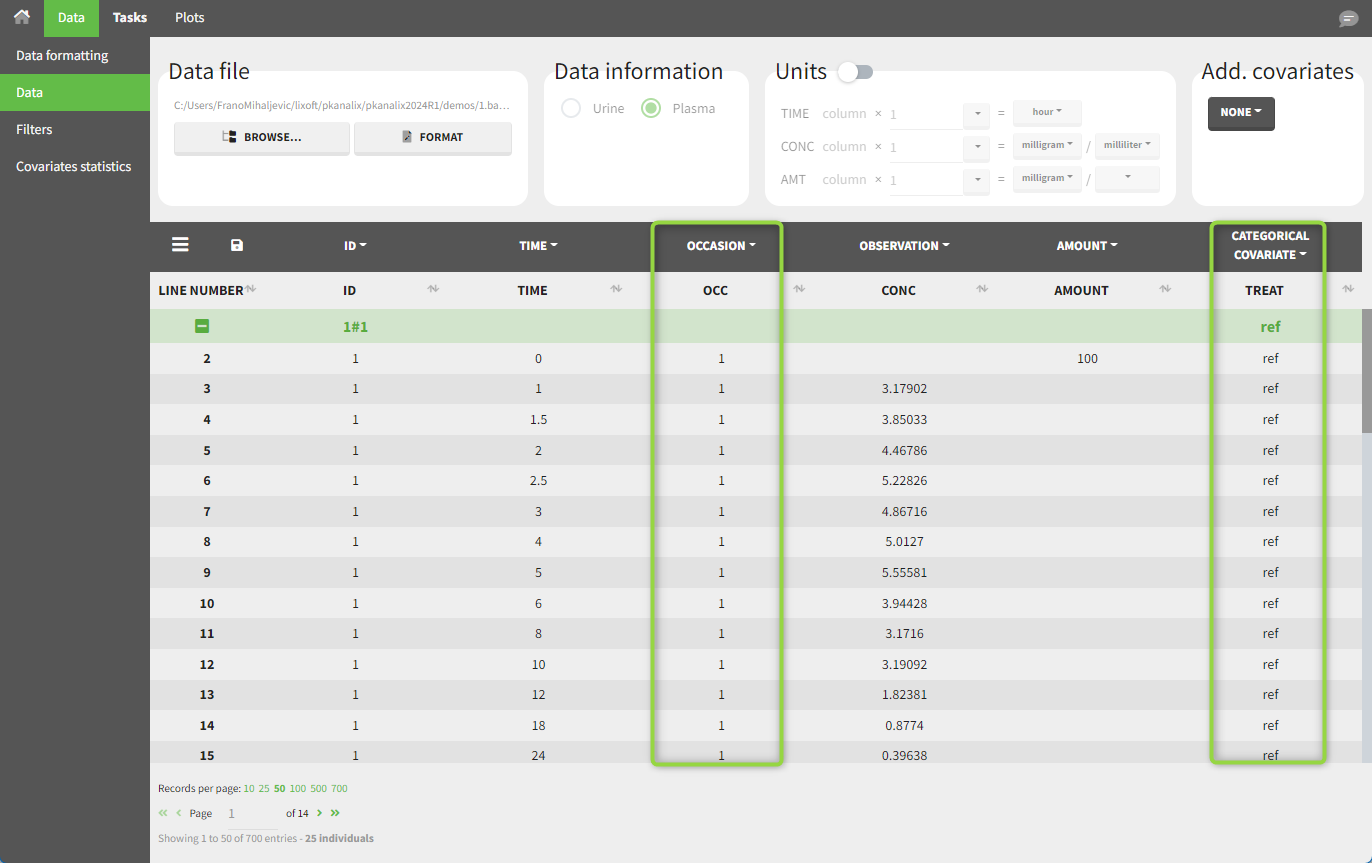
In the “Check lambda_z” tab (for the NCA) and the “Check init.” tab (for CA), each occasion for each individual is displayed. The syntax “1#2” indicates individual 1, occasion 2, according to the values defined in the ID and OCCASION columns.
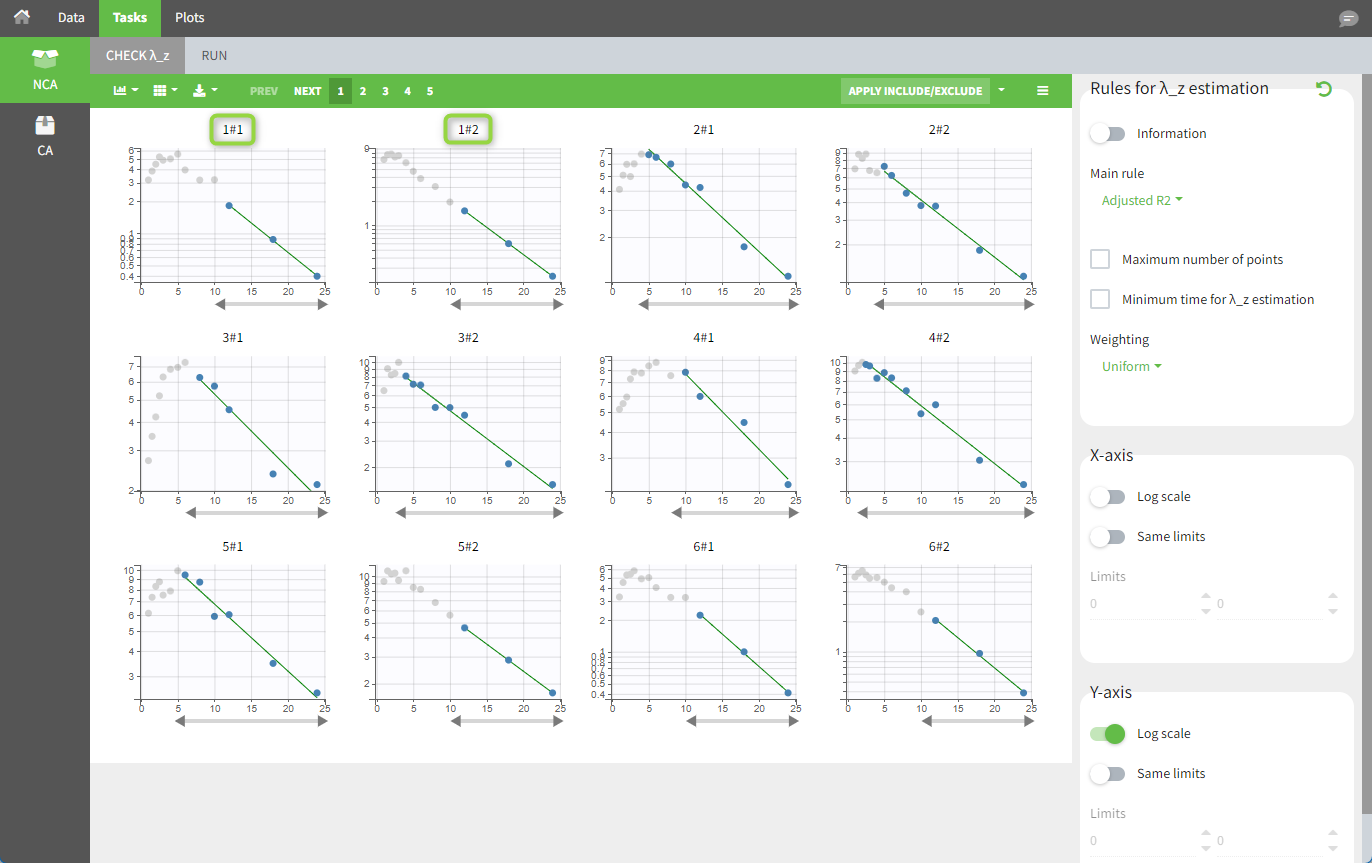
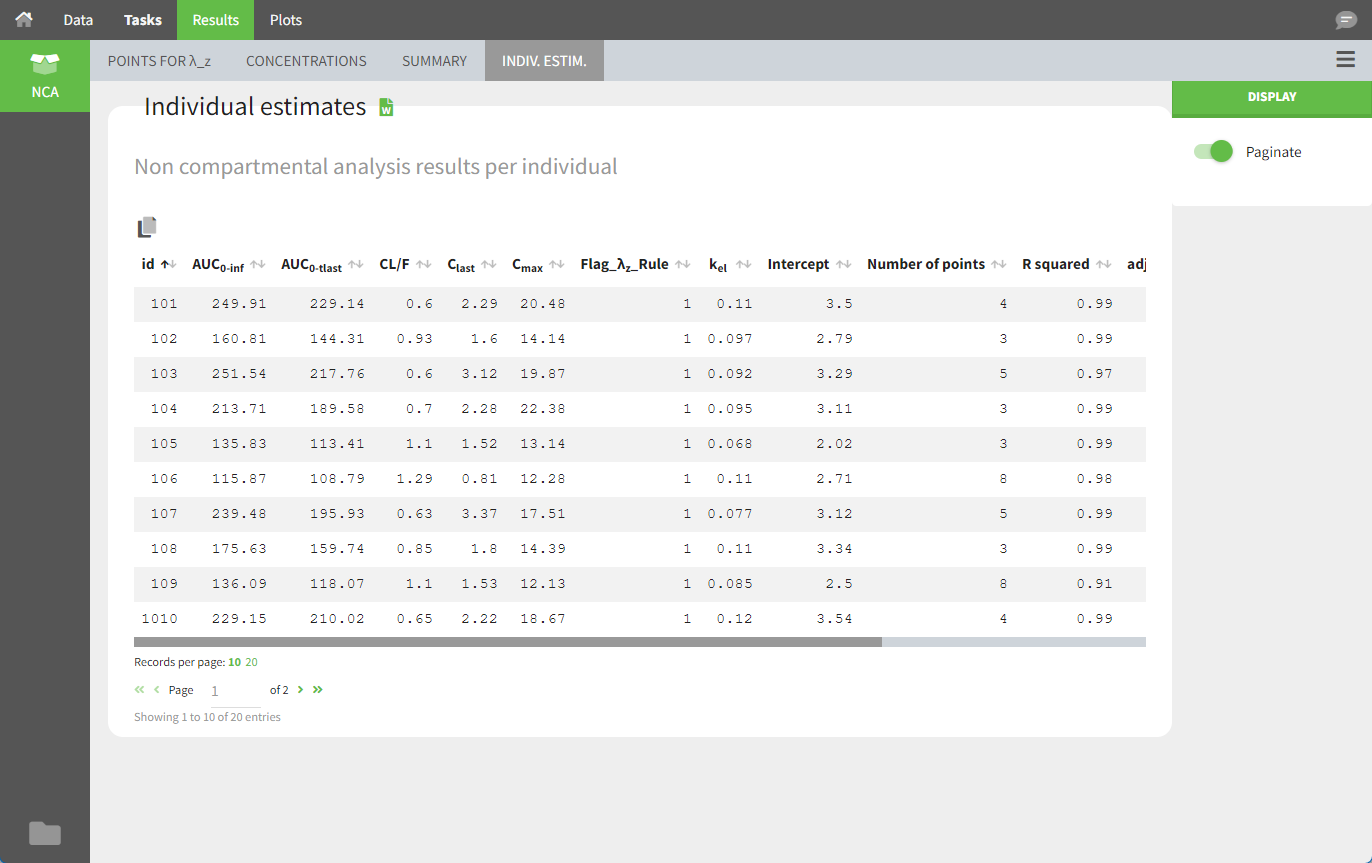
In the “Individual estimates” output tables, the first columns indicate the ID and OCCASION (using the dataset headers). The covariates are included at the end of the table. Note that it is possible to sort the table by any column, including ID, OCCASION and COVARIATES.
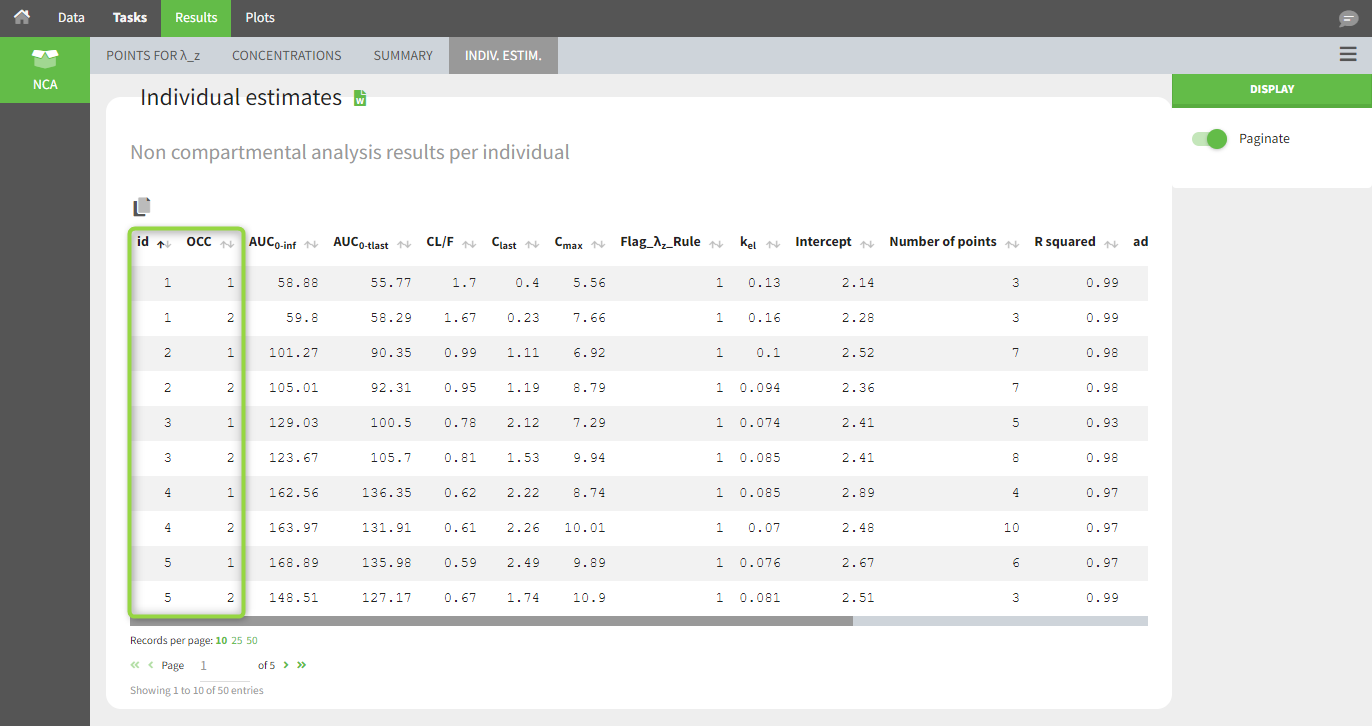
The OCCASION values (here “OCC”) are available to use in the plots for stratification, in addition to possible CATEGORICAL or CONTINUOUS COVARIATES (here “TREAT”).
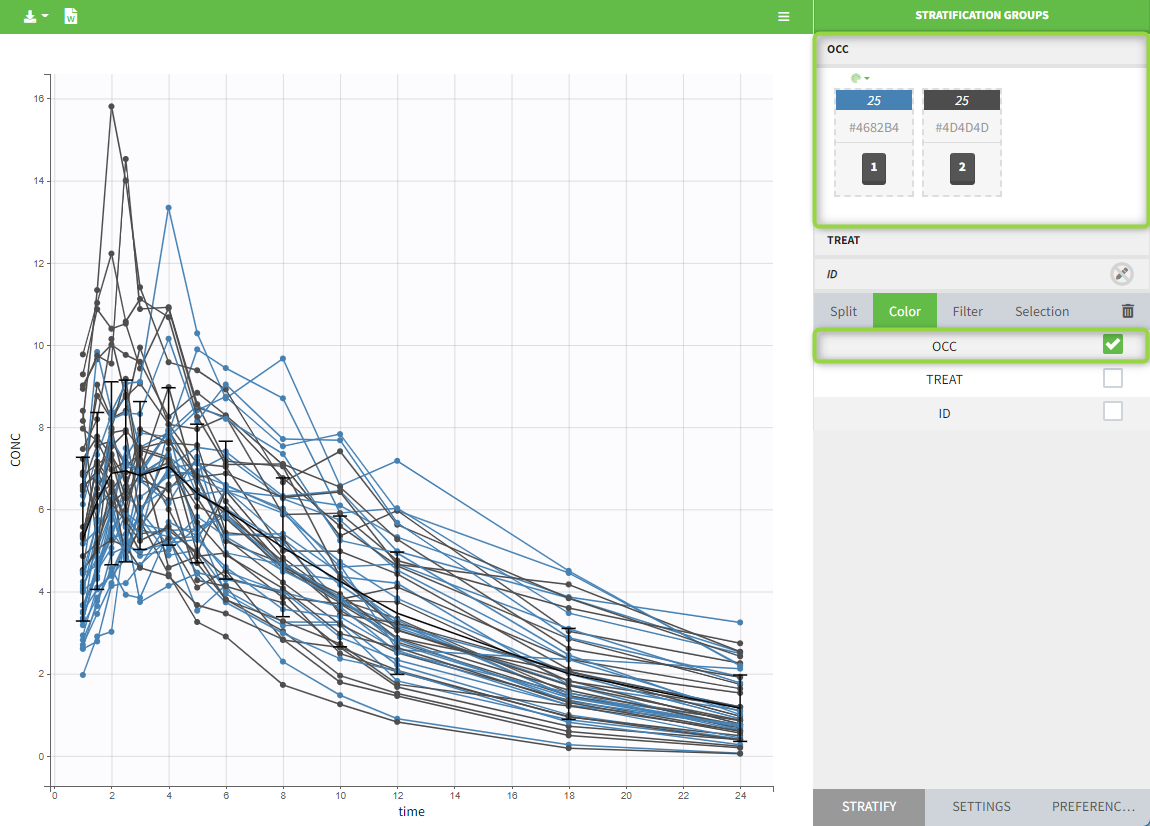
-
demo project_occasions2.pkx: a study with two treatments that are given with/without food
In this example, we have three sorting variables: ID, TREAT and FOOD. The TREAT and FOOD columns are duplicated: once with strings to be used as CATEGORICAL COVARIATE (TREAT and FOOD) and once with integers to be used as OCCASION (occT and occF).
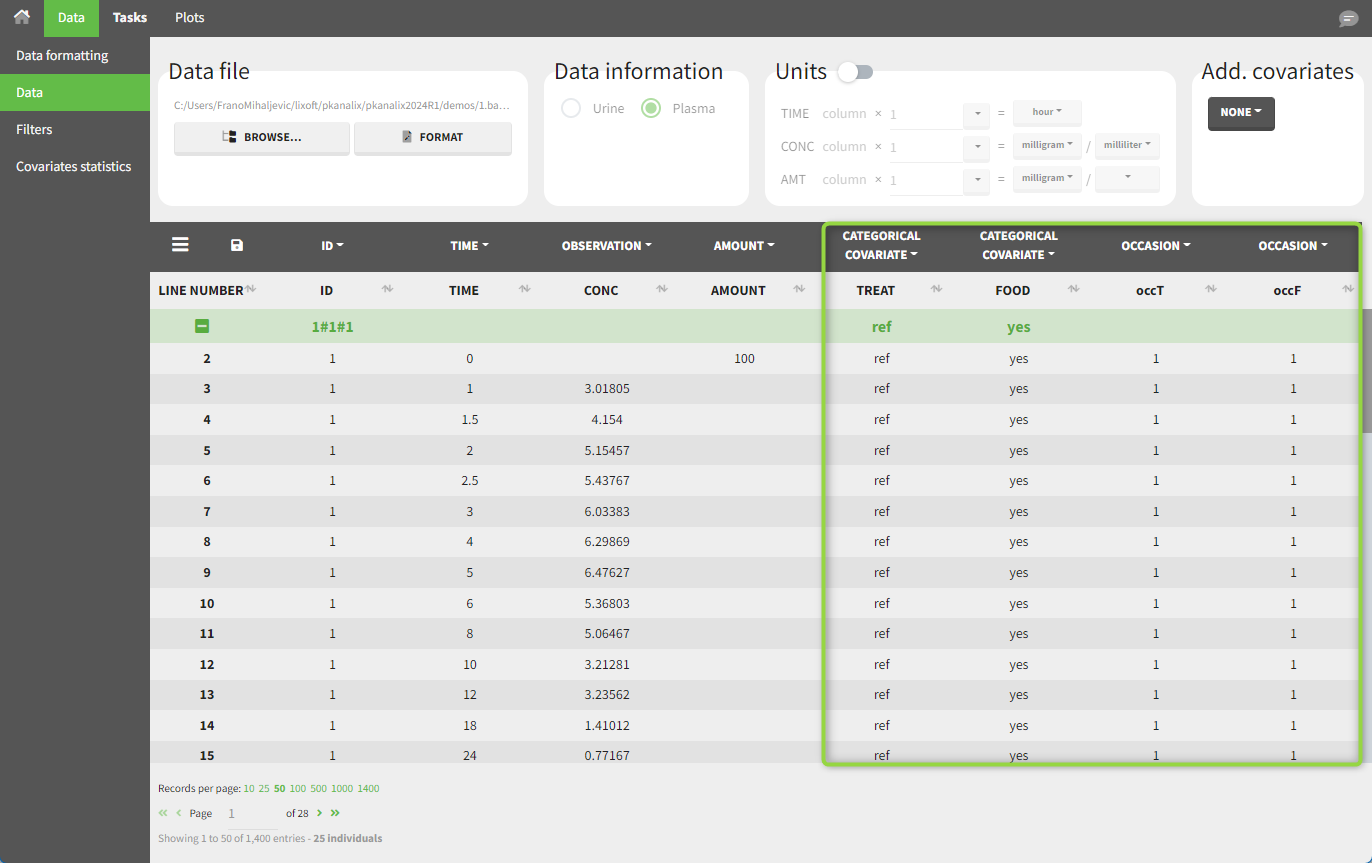
In the individual parameters tables and plots, three levels are visible (ID and the two occasions). In the “Individual parameters vs covariates” plot, we can plot Cmax versus FOOD, and split by TREAT for instance (Cmax versus TREAT and split by FOOD is also possible).
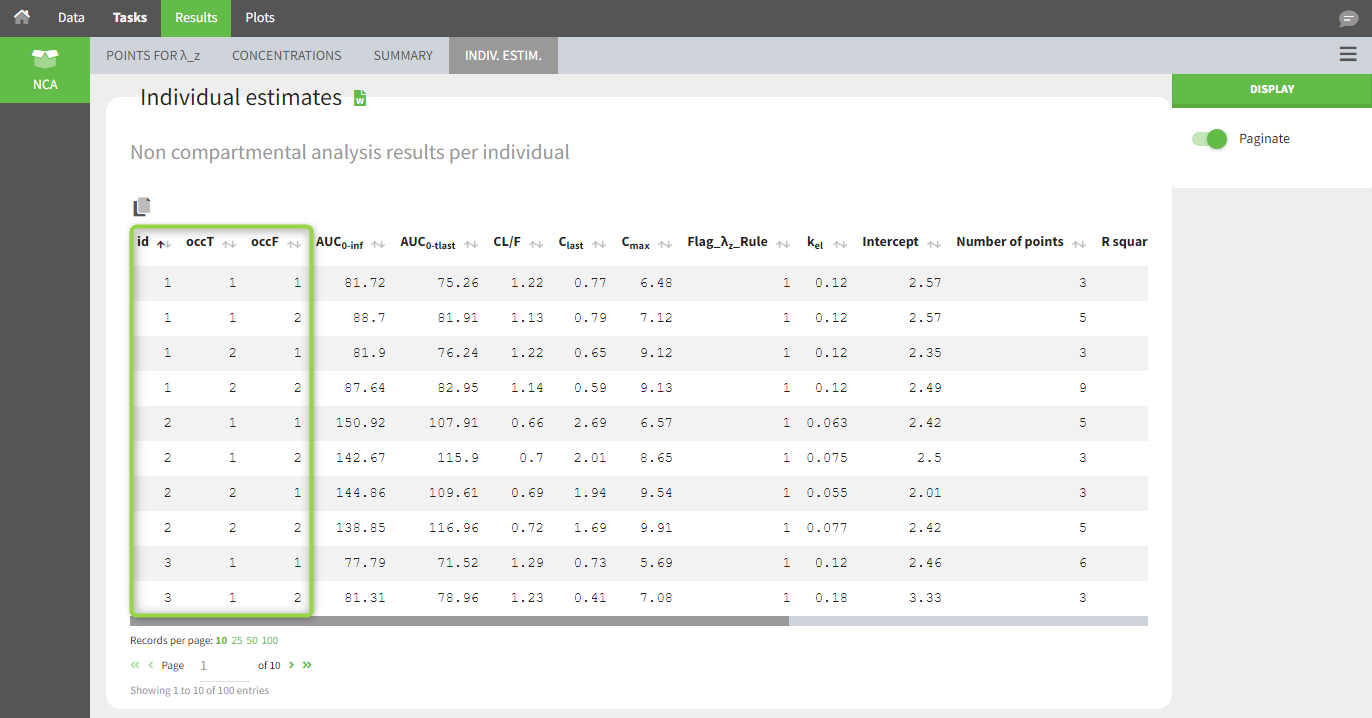
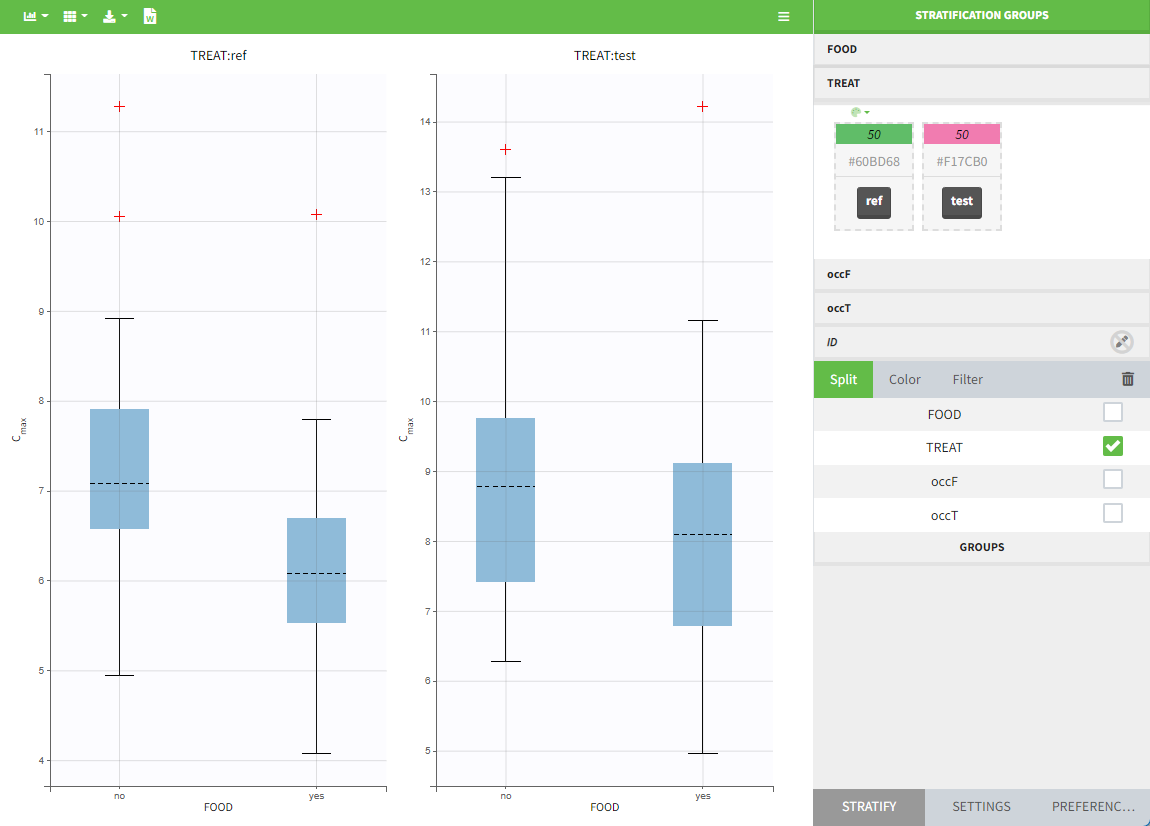
Covariates (“Carry” variables)
Individual information that should be carried over to output tables and plots must be tagged as CATEGORICAL or CONTINUOUS COVARIATES. Categorical covariates define variables with a few categories, such as treatment or sex, and are encoded as strings. Continuous covariates define variables on a continuous scale, such as weight or age, and are encoded as numbers. Covariates will not automatically be used as “Sort” variables. A dedicated OCCASION column is necessary (see above).
Covariates automatically appear in output tables. Plots of estimated NCA and/or CA parameters versus covariate values are also generated. In addition, covariates can be used to stratify (split, color or filter) plots. Statistics about the covariate distributions are available in table format in “Results > Cov. stat.” and in graphical format in “Plots > Covariate viewer”.
Avoid spaces and special characters (stars, etc.) in the strings for the categories of the categorical covariates. Underscores are allowed.
Example:
-
demo project_covariates.pkx
In this dataset, “SEX” is tagged as CATEGORICAL COVARIATE and “WEIGHT” as CONTINUOUS COVARIATE.
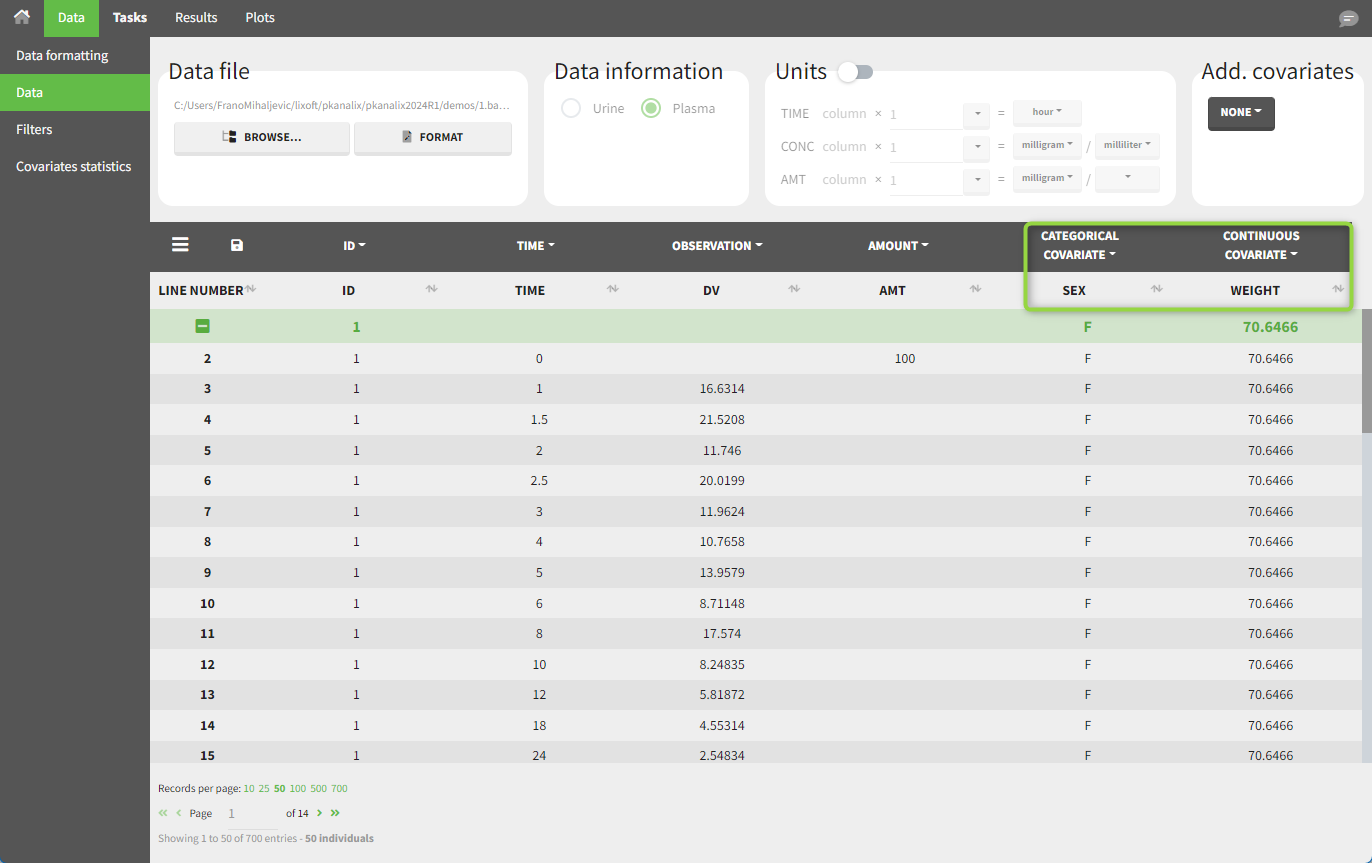
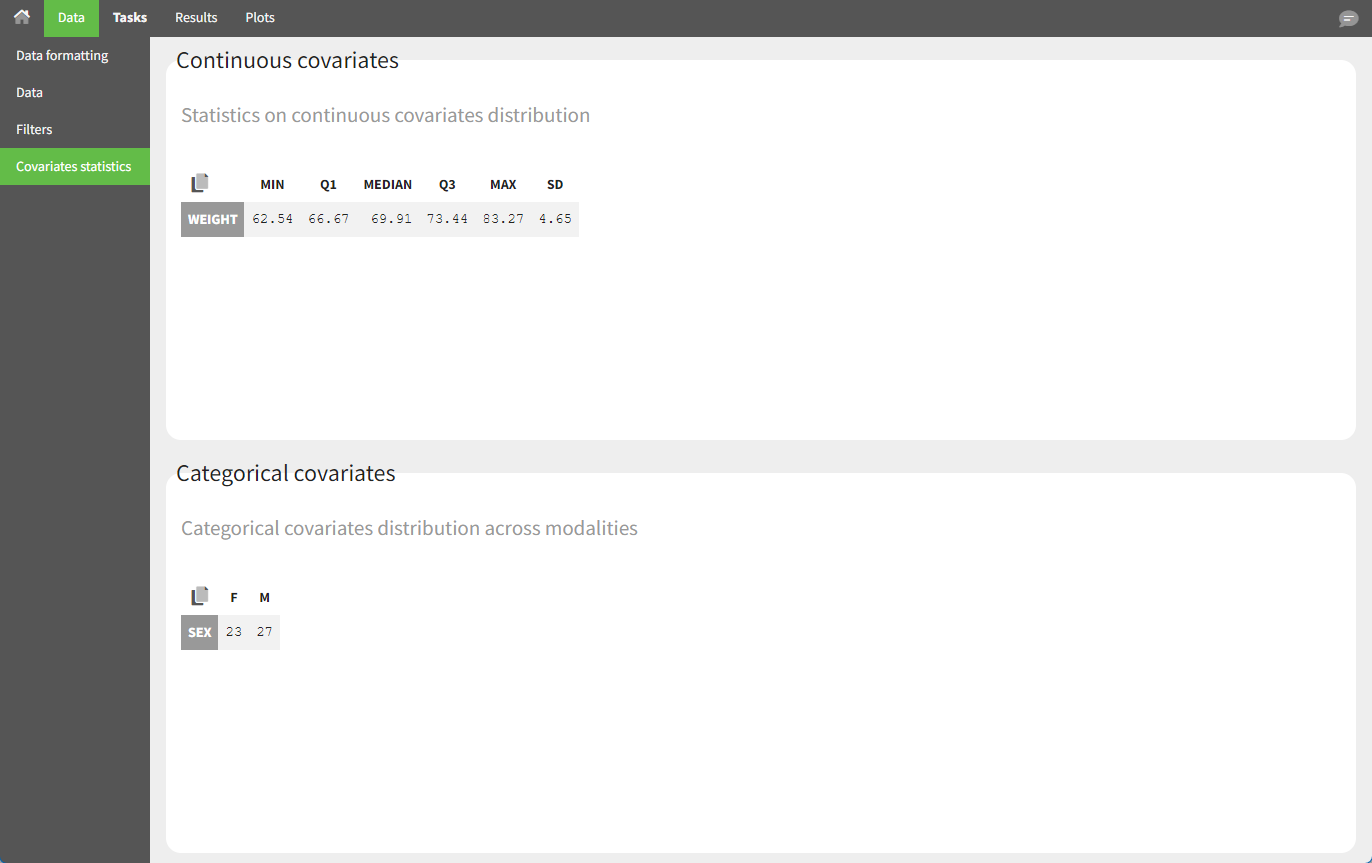
The “cov stat” table displays some statistics for the covariate values in the dataset. In the plot “Covariate viewer,” we see that the distribution of weight is similar for male and female subjects.
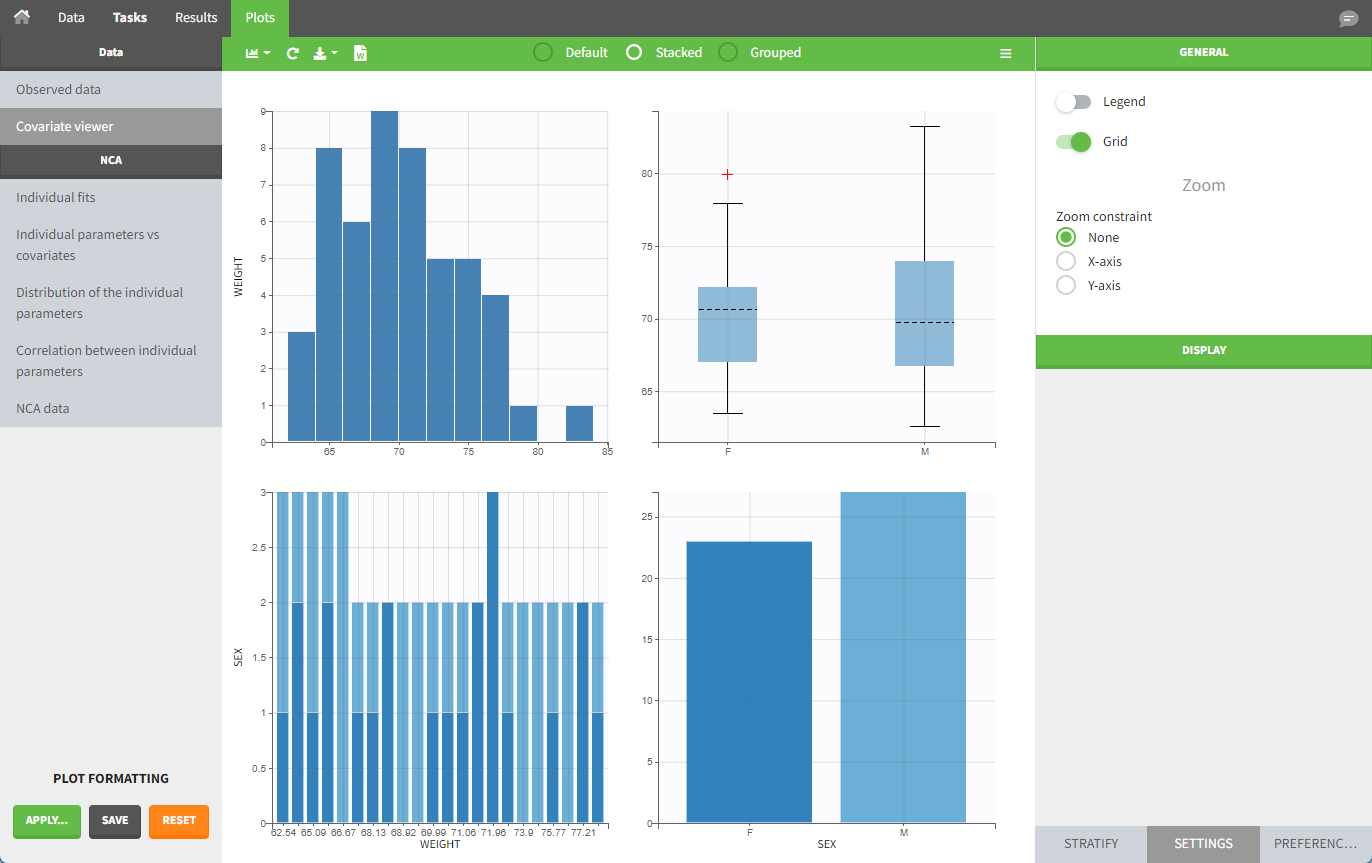
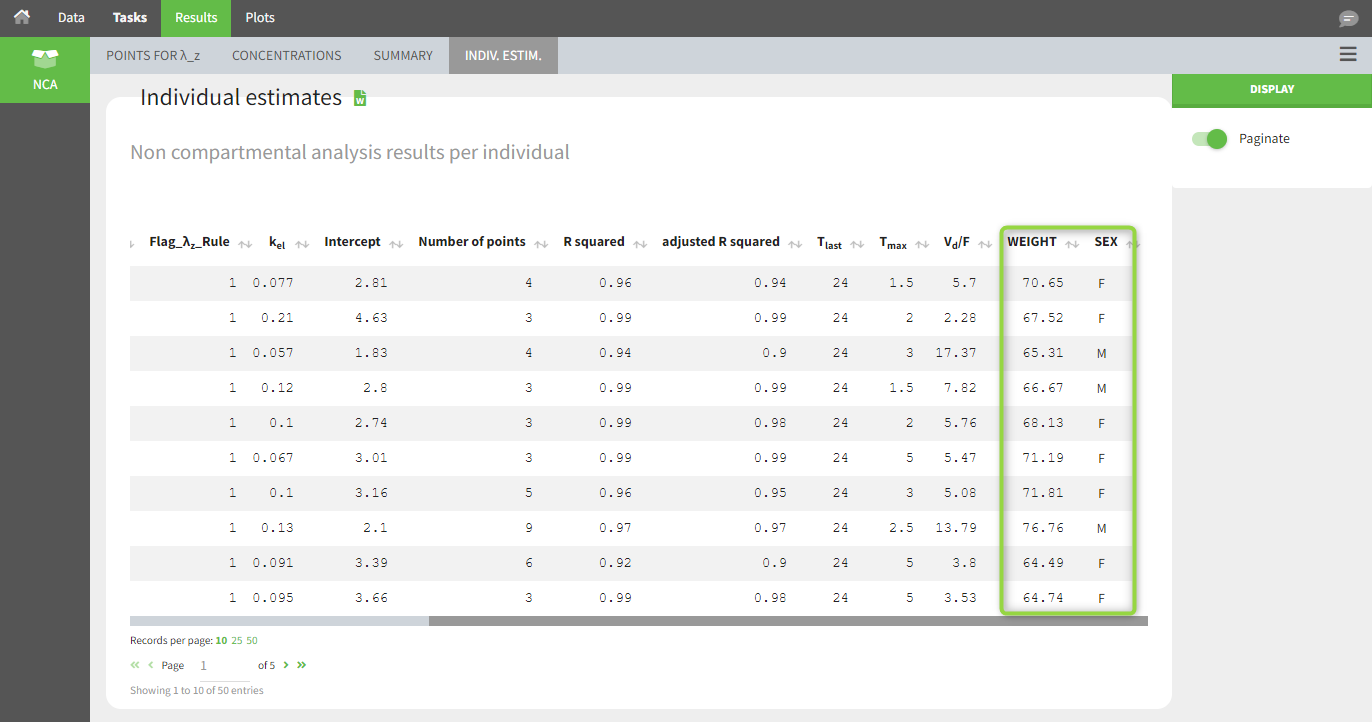
After running the NCA task, both covariates appear in the table of individual parameter estimates .
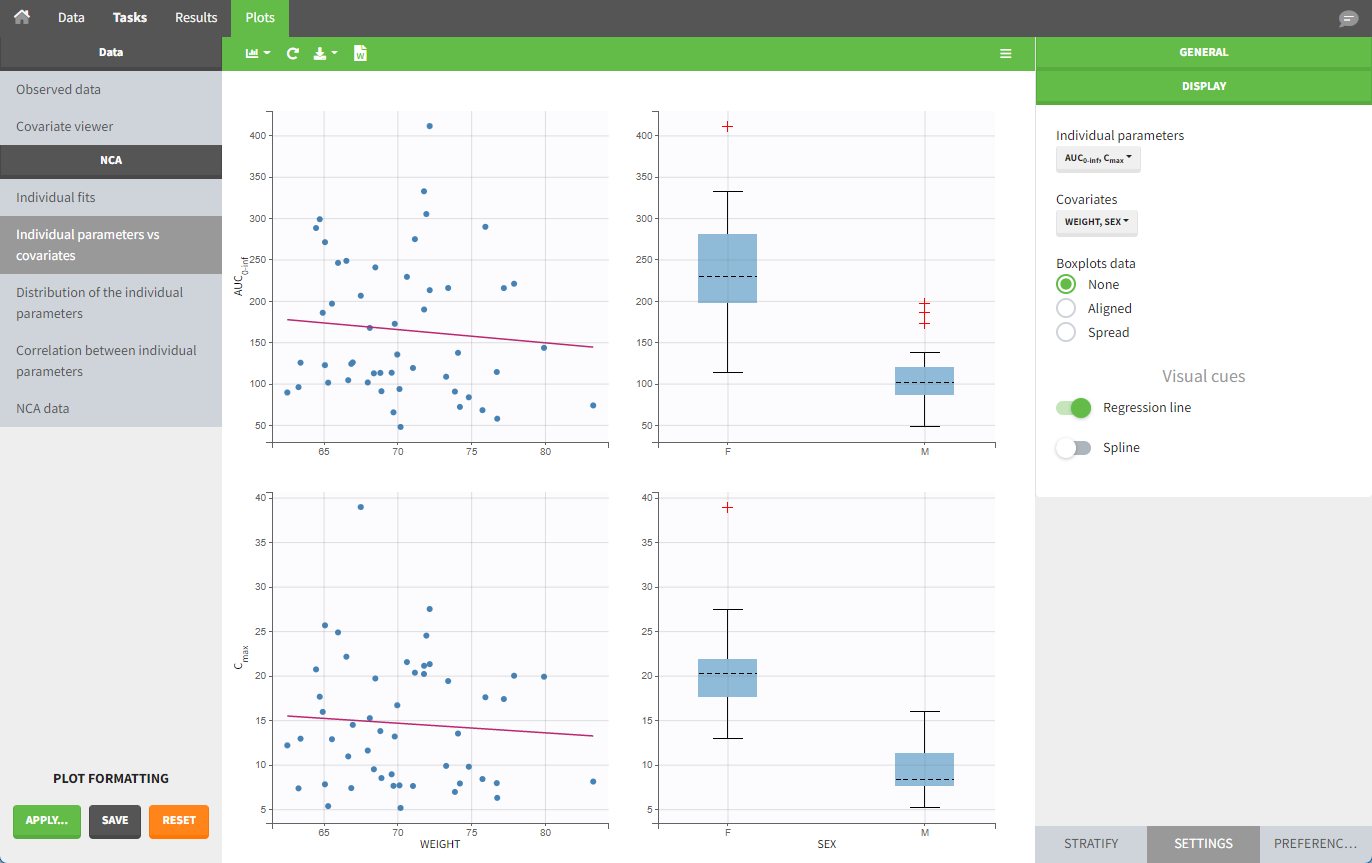
In the plot “parameters versus covariates,” the parameter values are plotted as scatter plots for continuous covariates with the parameter value (here Cmax and AUCINF_pred) on the y-axis and the covariate on the x-axis, and as boxplots for categorical covariates.
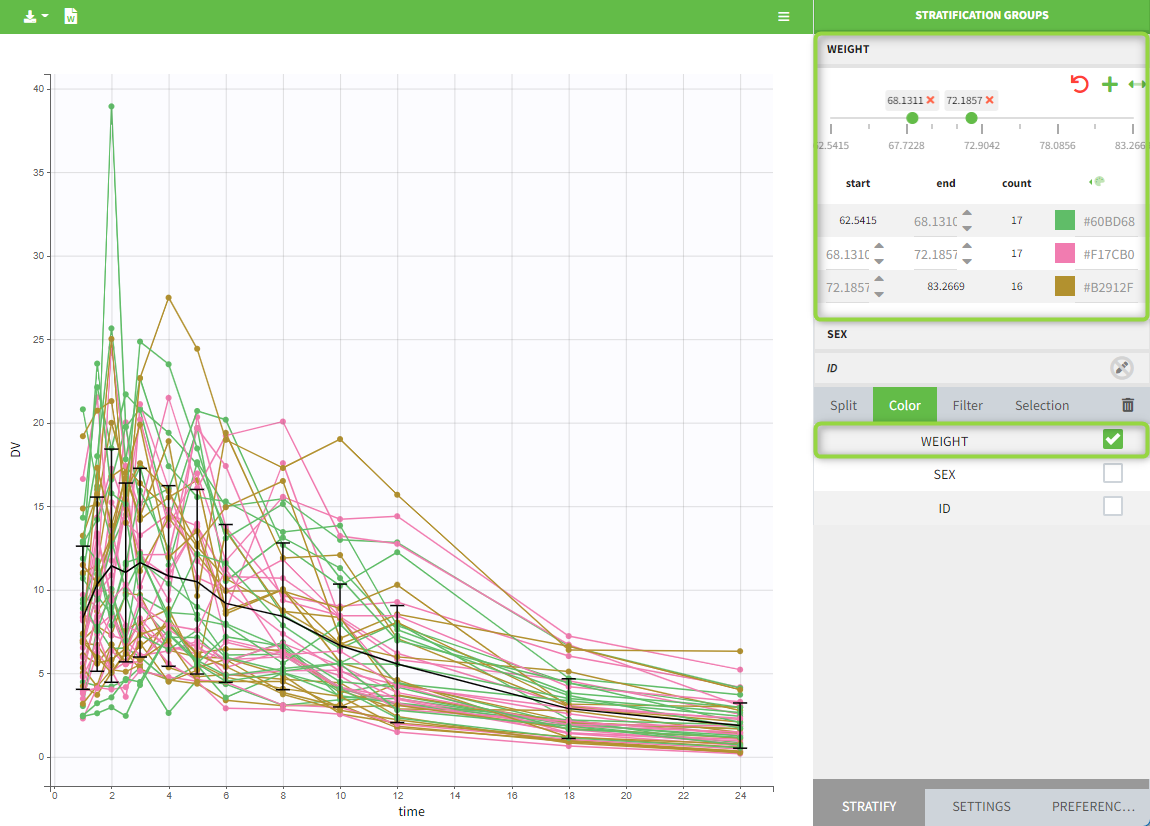
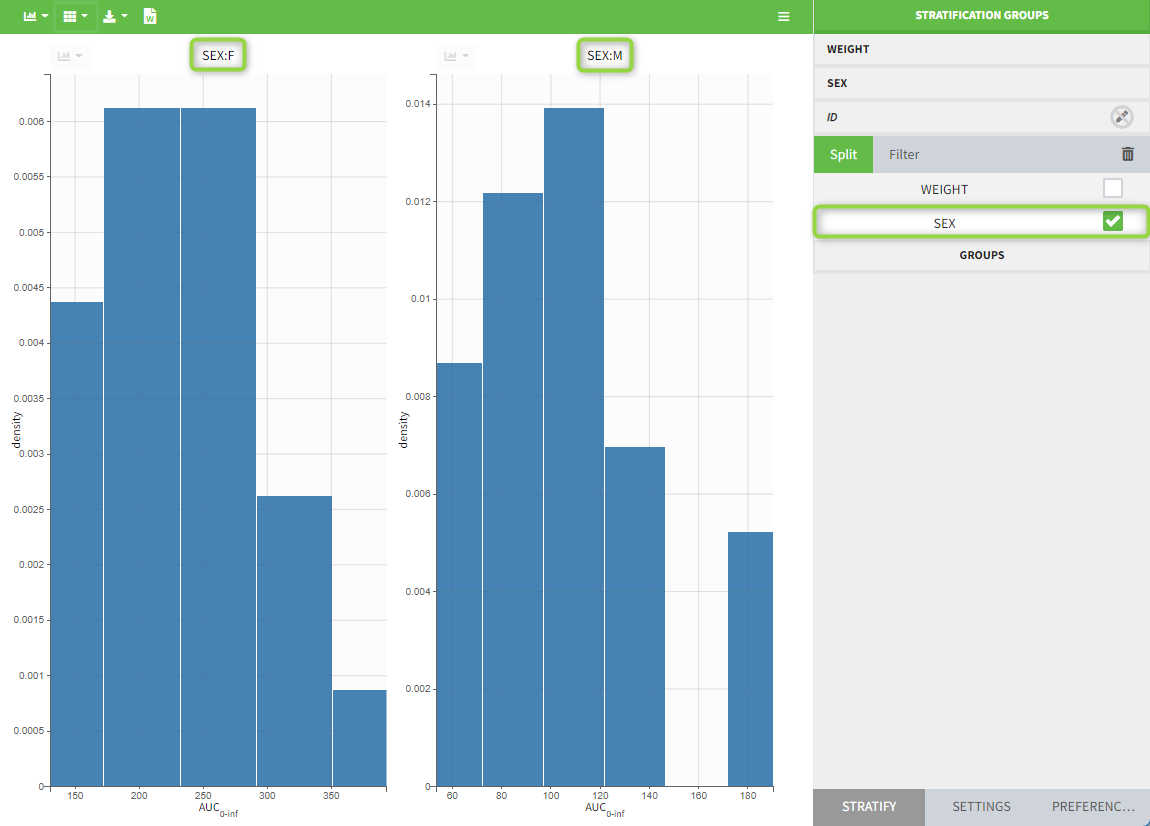
All plots can be stratified using the covariates. For instance, the “Observed data” plot can be colored by weight with three custom weight groups. Or the “Distribution of the parameters” plot can be split by sex, shown with the AUCINF_pred parameter.
Description of all possible column types
Column-types used for all types of lines:
-
OCCASION (formerly OCC): identifier (index) of the occasion
-
TIME (mandatory): time of the dose or observation record
-
NOMINAL TIME (from 2024 version): time at which doses and observations were expected to occur
-
DATE/DAT1/DAT2/DAT3: date of the dose or observation record, to be used in combination with the TIME column
-
EVENT ID (formerly EVID): identifier to indicate if the line is a dose-line or a response-line
-
IGNORED OBSERVATION (formerly MDV): identifier to ignore the OBSERVATION information of that line
-
IGNORED LINE (from 2019 version): identifier to ignore all the informations of that line
-
CONTINUOUS COVARIATE (formerly COV): continuous covariates (which can take values on a continuous scale)
-
CATEGORICAL COVARIATE (formerly CAT): categorical covariate (which can only take a finite number of values)
-
REGRESSOR (formerly X): defines a regression variable, i.e a variable that can be used in the structural model (used e.g for time-varying covariates)
-
IGNORE: ignores the information of that column for all lines
Column-types used for response-lines:
-
OBSERVATION (mandatory, formerly Y): records the measurement/observation for continuous, count, categorical or time-to-event data
-
OBSERVATION ID (formerly YTYPE): identifier for the observation type (to distinguish different types of observations, e.g PK and PD)
-
CENSORING (formerly CENS): marks censored data, below the lower limit or above the upper limit of quantification
-
LIMIT: upper or lower boundary for the censoring interval in case of CENSORING column
Column-types used for dose-lines:
-
AMOUNT (mandatory, formerly AMT): dose amount (with version 2024 only mandatory if dataset contains STEADY STATE, ADDITIONAL DOSES, INFUSIONRATE or INFUSION DURATION columns)
-
ADMINISTRATION ID (formerly ADM): identifier for the type of dose (given via different routes for instance)
-
INFUSION RATE (formerly RATE): rate of the dose administration (used in particular for infusions)
-
INFUSION DURATION (formerly TINF): duration of the dose administration (used in particular for infusions)
-
ADDITIONAL DOSES (formerly ADDL): number of doses to add in addition to the defined dose, at intervals INTERDOSE INTERVAL
-
INTERDOSE INTERVAL (formerly II): interdose interval for doses added using ADDITIONAL DOSES or STEADY-STATE column types
-
STEADY STATE (formerly SS): marks that steady-state has been achieved, and will add a predefined number of doses before the actual dose, at interval INTERDOSE INTERVAL, in order to achieve steady-state

